Non-Integrative mRNA Reprogramming: A Safer Pathway to Pluripotency for Regenerative Medicine and Drug Discovery
This article explores the transformative role of non-integrative mRNA technology in generating induced pluripotent stem cells (iPSCs).
Non-Integrative mRNA Reprogramming: A Safer Pathway to Pluripotency for Regenerative Medicine and Drug Discovery
Abstract
This article explores the transformative role of non-integrative mRNA technology in generating induced pluripotent stem cells (iPSCs). Aimed at researchers and drug development professionals, it details how transient mRNA delivery of reprogramming factors like OCT4, SOX2, KLF4, and c-Myc offers a safer alternative to genome-integrating methods by eliminating the risk of insertional mutagenesis. The content covers the foundational science, key delivery platforms such as synthetic mRNA and Sendai virus, and applications in disease modeling and personalized medicine. It also addresses critical challenges in efficiency, immunogenicity, and dosing control, while comparing the technology to other reprogramming approaches. The article concludes by evaluating the clinical translation of mRNA-derived iPSCs and future directions for this promising field.
The Science of Non-Integrative Reprogramming: Principles and Mechanisms of mRNA-Induced Pluripotency
The discovery of induced pluripotent stem cells (iPSCs) through the expression of specific transcription factors marked a revolutionary advance in regenerative medicine. However, the clinical translation of this technology has been hampered by the risks associated with genomic integration of foreign DNA. This review delineates the evolution from viral, gene-integrating methods to the development of non-integrative mRNA-based reprogramming technologies. We provide a comprehensive technical analysis of mRNA reprogramming methodologies, including synthetic modified mRNA and self-replicating RNA systems, detailing their underlying mechanisms, optimized protocols, and quantitative performance metrics. The transition to mRNA-based delivery represents a critical advancement toward generating clinically safe, footprint-free iPSCs for disease modeling, drug screening, and personalized cell-based therapies.
The seminal work of Takahashi and Yamanaka in 2006 demonstrated that somatic cells could be reprogrammed into induced pluripotent stem cells (iPSCs) through the forced expression of four transcription factors: OCT4, SOX2, KLF4, and c-Myc (OSKM) [1]. These Yamanaka factors activate a self-reinforcing pluripotency network, effectively reversing the developmental clock and granting somatic cells the capacity for unlimited self-renewal and differentiation into any cell type of the three germ layers [2].
Initial reprogramming methodologies relied on integrating viral vectors, particularly retroviruses and lentiviruses, which posed significant clinical risks due to their potential for insertional mutagenesis and tumorigenesis [3] [2]. The scientific community recognized that overcoming genomic integration was essential for clinical translation, sparking intensive research into non-integrative approaches including adenoviruses, plasmids, episomal vectors, and protein transduction [3].
Among these, mRNA-based reprogramming has emerged as a premier technology for clinical-grade iPSC generation. This approach offers an unambiguously footprint-free reprogramming method, as synthetic mRNA does not enter the nucleus or integrate into the host genome, and is rapidly degraded after translation [3] [4]. Subsequent advancements have addressed initial challenges of innate immune activation and transfection efficiency, establishing mRNA technology as a robust, safe, and highly efficient platform for cellular reprogramming in pluripotency research and regenerative medicine applications.
Evolution of Reprogramming Technologies
The progression from integrating to non-integrating reprogramming methods represents a critical pathway toward clinical applicability. The table below summarizes the key characteristics of major reprogramming vector systems.
Table 1: Comparison of Major Reprogramming Delivery Systems
| Vector Type | Genetic Material | Genomic Integration | Reprogramming Efficiency | Tumorigenic Risk | Primary Applications |
|---|---|---|---|---|---|
| Retrovirus | RNA | Yes (Random) | ~0.01% | High | Basic research |
| Lentivirus | RNA | Yes (Random) | ~0.1-1% | High | Basic research |
| Sendai Virus | RNA | No | ~0.1-1% | Low | Research & preclinical |
| Episomal Plasmid | DNA | Very Low (Random) | ~0.001% | Low | Research & preclinical |
| Protein Transduction | Protein | No | <0.001% | Very Low | Research |
| Modified mRNA | RNA | No | 1-4% | Very Low | Clinical translation |
| Self-replicating RNA | RNA | No | ~2-5% | Very Low | Clinical translation |
The Integration Problem
Initial retroviral and lentiviral systems for delivering OSKM factors provided the sustained transgene expression necessary for successful reprogramming but permanently altered the host cell genome [3]. This integration carries two significant risks: first, potential reactivation of silenced transgenes, some of which are known oncogenes (particularly c-Myc); and second, insertional mutagenesis through disruption of endogenous genes [2]. These safety concerns presented a substantial barrier to clinical translation, necessitating the development of non-integrative alternatives.
Key Non-Integrative Approaches
Early non-integrative methods included DNA-based plasmids and adenoviral vectors, but these systems typically showed reduced reprogramming efficiency due to transient gene expression and required repeated transfections [3]. Protein-based reprogramming represented another alternative but proved technically challenging with exceptionally low efficiency [3].
The emergence of RNA-based technologies provided a breakthrough, combining the safety of non-integration with high reprogramming efficiency. Two primary RNA platforms have been developed:
- Synthetic Modified mRNA: Incorporates nucleoside modifications (pseudouridine, 5-methylcytidine) to evade innate immune recognition while enabling robust, transient protein expression [4] [5].
- Self-replicating RNA (srRNA): Utilizes an alphavirus-derived replication system to amplify RNA copies within transfected cells, enabling prolonged transgene expression from a single transfection [4].
mRNA Reprogramming: Mechanisms and Methodologies
Molecular Basis of mRNA Reprogramming
The fundamental advantage of mRNA-based reprogramming lies in its cytoplasmic translation mechanism. Unlike DNA-based methods, mRNA does not require nuclear entry, and the translated reprogramming factors are produced as native proteins that readily localize to the nucleus to initiate pluripotency induction [4]. The transient nature of mRNA (typically degraded within 24-48 hours) necessitates repeated transfections but ensures no residual reprogramming activity persists once the process is complete.
A critical breakthrough in mRNA reprogramming came with the incorporation of nucleoside modifications (pseudouridine-Ψ and 5-methylcytidine) that dampen the innate immune response by reducing recognition by pattern recognition receptors [4] [6]. Additionally, optimized 5' cap structures (Cap1) and elongated poly(A) tails significantly enhance translation efficiency and mRNA stability [4] [7].
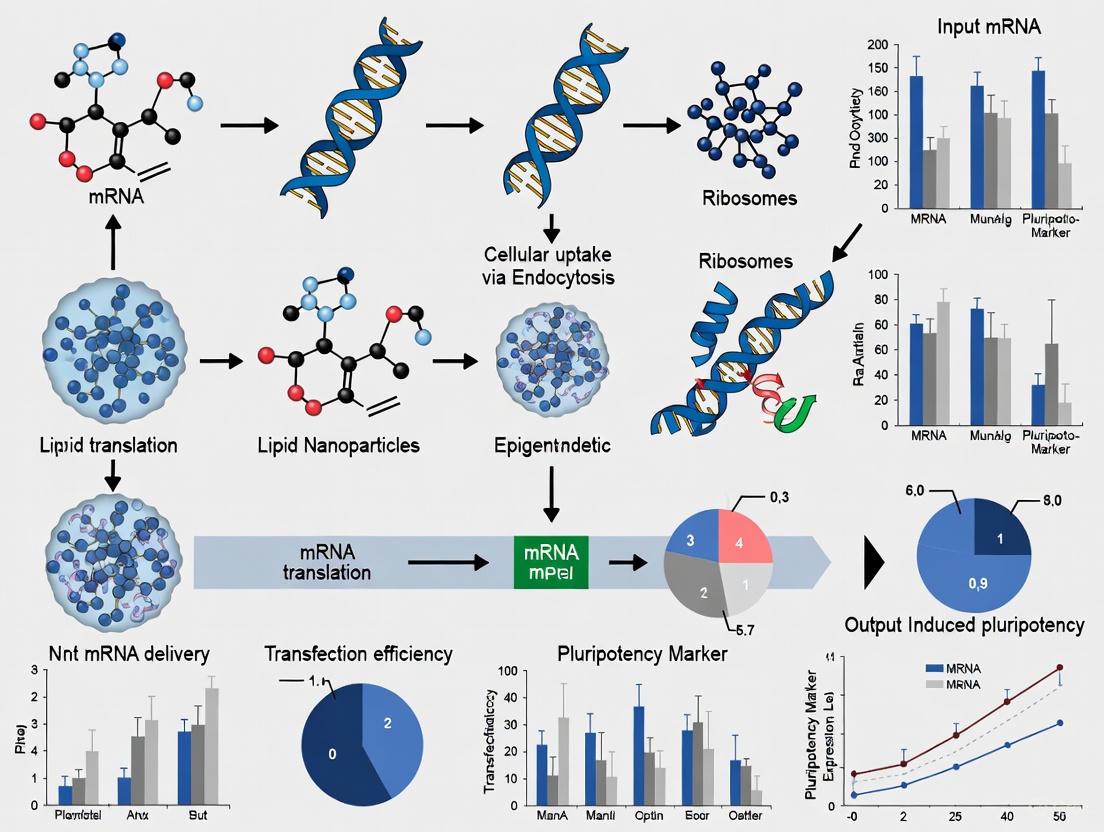
Diagram 1: mRNA Reprogramming Workflow and Mechanism
Quantitative Performance of mRNA Systems
Comparative studies have demonstrated significant differences in performance between conventional synthetic mRNA and self-replicating RNA systems. The table below summarizes key efficiency metrics from direct comparative studies.
Table 2: Performance Metrics of mRNA Reprogramming Systems
| Parameter | Standard Modified mRNA | Self-replicating RNA (srRNA) | Reference |
|---|---|---|---|
| Transfection Frequency | Daily (14-18 transfections) | Single transfection | [4] |
| Total RNA Required | ~1.2μg daily (≥16.8μg total) | 1μg single dose | [4] |
| Reprogramming Efficiency | 1-4% | ~2-5% | [4] [6] |
| Time to Colony Emergence | 14-24 days | 10-18 days | [4] |
| Immune Suppression Required | B18R essential | B18R essential | [4] |
| GFP Reporter Tracking | Not available | Enabled by IRES-GFP | [4] |
Detailed Experimental Protocol: mRNA Reprogramming
This section provides a comprehensive methodology for generating iPSCs using synthetic modified mRNA, based on established protocols [4] [7] [6].
mRNA Synthesis and Preparation
- Template Design: Generate DNA templates containing the coding sequences for human OCT4, SOX2, KLF4, c-MYC, and optionally LIN28 using pcDNA3.3-based plasmids. Include a T120 sequence for poly(A) tail addition.
- In Vitro Transcription (IVT): Perform IVT reactions using:
- 1.5μg linearized DNA template
- ATP, GTP, pseudouridine-5'-triphosphate (ΨTP), and 5-methylcytidine-5'-triphosphate (5mCTP)
- 3'-0-Me-m7G(5')ppp(5')G RNA Cap Structure Analog or enzymatic capping
- Incubate at 37°C for 4 hours
- Post-Transcriptional Modifications:
- Enzymatic 5' capping using ScriptCap Cap1 Capping System
- 3' polyadenylation using A-Plus Poly(A) Polymerase Tailing Kit
- Purification and Quality Control: Purify mRNA using silica membrane-based kits. Verify integrity by formaldehyde-agarose gel electrophoresis and quantify by spectrophotometry. Adjust concentration to 100ng/μL in nuclease-free water.
Cell Culture and Transfection
Somatic Cell Preparation:
- Culture neonatal human foreskin fibroblasts (NuFFs) in DMEM high glucose supplemented with 10% FBS, 1× GlutaMAX, and 50μg/mL gentamicin.
- Seed fibroblasts at 5×10⁴ cells per well of a 6-well plate 24 hours before transfection.
Transfection Protocol:
- Prepare mRNA-lipid complexes using 1.2μg mRNA mix and appropriate transfection reagent per manufacturer's instructions.
- Replace cell culture medium with fresh medium containing B18R protein (0.25μg/mL) to suppress interferon response.
- Add mRNA-transfection complexes to cells.
- Repeat transfection daily for 16-18 days.
iPSC Colony Selection and Expansion:
- Monitor for emergence of compact, ES-like colonies from day 10 onward.
- Pick established colonies manually and transfer to mitotically inactivated mouse embryonic fibroblast (MEF) feeders in iPSC culture medium.
- Expand and characterize clonal lines through immunocytochemistry (OCT4, NANOG, SSEA-4), teratoma formation, and pluripotency gene expression analysis.
The Scientist's Toolkit: Essential Reagents for mRNA Reprogramming
Successful implementation of mRNA reprogramming requires carefully selected reagents and components. The following table details essential research reagent solutions for establishing this technology.
Table 3: Essential Research Reagents for mRNA Reprogramming
| Reagent Category | Specific Product/Component | Function and Application Notes |
|---|---|---|
| Reprogramming Factors | Synthetic mRNA encoding OCT4, SOX2, KLF4, c-MYC | Core reprogramming factors; modified nucleosides (Ψ, 5mC) essential for immune evasion |
| Immune Suppression | B18R recombinant protein | Interferon inhibitor; critical for cell viability during repeated transfections |
| Transfection Reagent | Lipofectamine RNAiMAX or polyethylenimine (PEI) | Facilitates cellular uptake of mRNA; PEI offers cost-effective alternative |
| Somatic Cell Source | Neonatal human foreskin fibroblasts (NuFFs) | Well-characterized starting cell type; alternative sources include peripheral blood mononuclear cells |
| Culture Medium | DMEM high glucose + 10% FBS (fibroblasts); Essential 8 Medium (iPSCs) | Supports somatic cell growth and pluripotent stem cell maintenance |
| Feeder Cells | Mitomycin C-treated MEFs or NuFFs | Provides extracellular matrix and secreted factors supporting iPSC colony formation |
| Quality Control Assays | Pluripotency markers (OCT4, NANOG, SSEA-4); Karyotyping; Trilineage differentiation | Validates successful reprogramming and genomic integrity |
Applications and Future Directions
The transition to mRNA-based reprogramming has significantly advanced the clinical translation of iPSC technology. Key application areas include:
Disease Modeling and Drug Screening
Patient-specific iPSCs generated via mRNA reprogramming provide robust platforms for investigating disease mechanisms and screening therapeutic compounds. This approach has been particularly valuable for neurodegenerative diseases such as amyotrophic lateral sclerosis (ALS), where iPSC-derived motor neurons recapitulate disease-specific pathology [1]. Similar strategies are being applied to model progeroid syndromes and other genetic disorders [8].
Regenerative Medicine and Cell Therapy
The non-integrating nature of mRNA-reprogrammed iPSCs makes them ideal candidates for cell-based therapies. Directed differentiation of these iPSCs can generate autologous cells for transplantation, including dopaminergic neurons for Parkinson's disease, cardiac cells for myocardial repair, and pancreatic beta cells for diabetes [9] [6].
Partial Reprogramming for Rejuvenation
Recent advances have explored partial reprogramming through transient mRNA expression of Yamanaka factors to reverse age-associated cellular changes without complete dedifferentiation. This approach has demonstrated potential for rejuvenating aged cells and tissues, restoring function in mouse models of aging and disease [10] [8].
Diagram 2: Applications of mRNA Reprogramming Technology
The evolution from Yamanaka's original viral factors to contemporary mRNA-based reprogramming represents a transformative advancement in cellular engineering. This transition has effectively addressed the critical safety concerns associated with genomic integration while achieving superior reprogramming efficiencies. The development of nucleoside-modified mRNA and self-replicating RNA systems has enabled robust, footprint-free generation of iPSCs suitable for clinical applications.
Current mRNA reprogramming protocols provide researchers with powerful tools for disease modeling, drug discovery, and regenerative medicine. The continued refinement of mRNA design, delivery methods, and differentiation protocols will further enhance the clinical potential of this technology. As the field progresses, mRNA-based reprogramming is poised to become the gold standard for generating clinically relevant iPSCs, ultimately enabling personalized regenerative therapies for a broad spectrum of human diseases.
Transient mRNA expression has emerged as a powerful non-integrative technology for reprogramming somatic cells, offering unprecedented control over epigenetic and transcriptional resetting. This technical guide explores the core mechanisms by which short-lived mRNA transcripts of reprogramming factors can orchestrate a profound reconfiguration of the cellular epigenome. Unlike integrating vector systems, transient mRNA delivery achieves reprogramming without permanent genetic alteration, making it particularly valuable for both basic pluripotency research and therapeutic applications. We examine the molecular dynamics of this process, including the rapid erasure of somatic epigenetic memory, the establishment of youthful transcriptional networks, and the resetting of epigenetic clocks. Through structured data presentation, detailed protocols, and pathway visualizations, this review provides researchers with a comprehensive toolkit for implementing and advancing this transformative technology.
The discovery that somatic cells can be reprogrammed to induced pluripotent stem cells (iPSCs) through exogenous expression of specific transcription factors revolutionized regenerative medicine and disease modeling [11]. Initial reprogramming methods relied on integrating viral vectors, which posed significant safety concerns for clinical applications due to the risk of insertional mutagenesis and persistent transgene expression [4]. Transient mRNA-based technology has overcome these limitations by delivering reprogramming factors as synthetic mRNA molecules that are translated into proteins but do not integrate into the host genome [4] [9]. This non-integrative approach provides precise temporal control over factor expression and eliminates the risk of genomic alteration, making it particularly suitable for generating clinical-grade iPSCs [4].
The core advantage of transient mRNA expression lies in its ability to reset epigenetic and transcriptional networks without permanent genetic modification. After delivery into the cytosol, the mRNA is immediately translated by ribosomes into reprogramming proteins, and the synthesis of these factors ceases once the mRNA degrades, leaving no footprints in the genome [4]. This transient expression profile is sufficient to initiate a cascade of epigenetic remodeling events that ultimately lead to the acquisition of pluripotency [12]. The technology has been further refined through nucleoside modifications that enhance mRNA stability and reduce innate immune recognition, alongside the use of interferon inhibitors to prevent cellular stress responses during the reprogramming process [4].
Molecular Mechanisms of Epigenetic and Transcriptional Resetting
Dynamics of Epigenomic Remodeling
Transient mRNA reprogramming initiates a profound reconfiguration of the epigenetic landscape through coordinated mechanisms. During early reprogramming phases, exogenous mRNA-derived OSKMLN (OCT4, SOX2, KLF4, c-MYC, LIN28, NANOG) factors bind to somatic enhancers, recruiting chromatin remodelers away from loci maintaining somatic identity [13]. This binding sequesters transcription factors like AP-1 (JUN/FOS) from somatic enhancers, initiating their silencing while simultaneously activating early pluripotency networks [13]. The process involves two distinct transcriptional waves: early stochastic events where somatic genes are silenced and early pluripotency genes activated, followed by more deterministic late-phase events where late pluripotency-associated genes are established [11].
DNA methylation changes follow a specific temporal sequence during transient reprogramming. Analysis throughout primed and naive reprogramming reveals that aberrant DNA methylation differences characteristic of conventional iPSCs emerge midway through primed reprogramming (between days 13-21), whereas DNA demethylation begins early in naive reprogramming [13]. The transient expression approach enables resetting of the epigenetic clock without reaching the point of no return in complete reprogramming, allowing cells to maintain their identity while reversing age-associated epigenetic marks [12]. This partial reprogramming strategy targets age-related epigenetic changes without completely erasing cellular identity, making it particularly valuable for therapeutic applications aimed at reversing cellular aging while retaining tissue-specific function [12].
Transcriptional Network Resetting
Transient mRNA expression rapidly activates more youthful gene expression profiles without affecting cell identity genes. In aged human fibroblasts and endothelial cells, just four days of OSKMLN mRNA transfection followed by a two-day interruption was sufficient to shift transcriptional profiles toward younger patterns [12]. Bulk RNA sequencing revealed that 24.7% of genes differentially expressed between young and aged fibroblasts overlapped with genes changed by transient reprogramming, with the directionality of change matching that of youth [12]. This rejuvenation effect occurred without detectable expression of pluripotency markers beyond the transfected mRNAs and without significant changes to established cell identity markers [12].
The transcriptional resetting extends to critical aging pathways, including reduction of inflammatory profiles in aged chondrocytes and restoration of youthful regenerative responses in aged human muscle stem cells [12]. The process reactivates autophagy and proteasomal activity pathways that typically decline with age, enhancing cellular proteostasis [12]. The interspecies mRNA transfer research further demonstrates that transferred reprogramming factor mRNAs (including Tfcp2l1, Tfap2c, and Klf4) are translated into functional proteins that directly contribute to altering the pluripotency state in acceptor cells [14]. This confirms that transiently expressed mRNAs can directly impact transcriptional networks without genomic integration.
Quantitative Data on Epigenetic and Transcriptional Changes
Epigenetic Clock Reversal and Gene Expression Changes
Table 1: Epigenetic Clock Reversal Following Transient mRNA Reprogramming
| Cell Type | Epigenetic Clock | Average Age Reversal (Years) | Standard Error | Statistical Significance |
|---|---|---|---|---|
| Endothelial Cells | Horvath pan-tissue | -4.94 | 1.63 | P = 0.023 |
| Fibroblasts | Horvath pan-tissue | -1.84 | 1.46 | P = 0.023 |
| Combined Cells | Horvath pan-tissue | -3.40 | 1.17 | P = 0.023 |
| Endothelial Cells | Skin-and-blood clock | -1.62 | 0.67 | P = 0.042 |
| Fibroblasts | Skin-and-blood clock | -1.07 | 0.67 | P = 0.042 |
Table 2: Transcriptional Changes in Aged Human Cells After Transient Reprogramming
| Cell Type | Total Genes Changed | Upregulated Genes | Downregulated Genes | Overlap with Youthful Signature |
|---|---|---|---|---|
| Fibroblasts | 1,042 | 734 | 308 | 24.7% (odds ratio 4.53) |
| Endothelial Cells | 992 | 461 | 531 | 16.7% (odds ratio 3.84) |
Table 3: Reprogramming Efficiency Comparison Between mRNA Methods
| Method | Reprogramming Time | RNA Amount | Key Advantages | Efficiency |
|---|---|---|---|---|
| Synthetic mRNA | Daily transfection for 14+ days | 1.2μg per day | No genomic integration; immediate translation | Standard efficiency |
| Self-replicating RNA (srRNA) | Single transfection | 1μg single dose | Extended protein expression; more efficient | Significantly improved |
Hallmarks of Aging Amelioration
Transient mRNA reprogramming produces measurable improvements across multiple cellular hallmarks of aging. In aged human fibroblasts and endothelial cells, treatment increased nuclear levels of the epigenetic repressive mark H3K9me3, heterochromatin-associated protein HP1γ, and nuclear lamina support protein LAP2α, all of which typically decrease with age [12]. The reprogramming also enhanced proteostatic mechanisms, increasing both autophagosome formation and chymotrypsin-like proteasomal activity in aged cells [12]. These improvements occurred without abolishing cellular identity, as verified through retention of cell-type-specific markers and absence of pluripotency marker activation beyond the transfected factors [12].
The technology demonstrates particular efficacy in resetting epigenetic memory concentrated in cell-of-origin-dependent repressive chromatin marked by H3K9me3, lamin-B1, and aberrant CpH methylation [13]. Transient naive reprogramming reconfigures these domains to an embryonic stem cell-like state without disrupting genomic imprinting, effectively correcting the transposable element overexpression and differential gene expression seen in conventional iPSCs [13]. The resulting cells show similar differentiation efficiencies to embryonic stem cells, addressing a major limitation of conventional iPSC generation methods [13].
Experimental Protocols for Transient mRNA Reprogramming
mRNA Synthesis and Preparation
Synthetic mRNA Synthesis Protocol:
- DNA Template Generation: Use pcDNA 3.3 plasmids containing coding sequences for Klf4, cMyc, Oct4, Sox2, Lin28, or eGFP. Generate DNA templates via PCR using forward primer (5′-TTGGACCCTCGTACAGAAGCTAATACG-3′) and reverse primer (5′-T120CTTCCTACTCAGGCTTTATTCAAAGACCA-3′) to incorporate a polyT120 sequence [4].
In Vitro Transcription (IVT): Perform IVT reaction with 1.5μg DNA template, ATP, GTP, pseudouridine-5′-triphosphate (Pseudo-UTP), 5-methylcytidine-5′-triphosphate (5mCTP), and 3′-0-Me-m7G(5′)ppp(5′)G RNA Cap Structure Analog. Incubate at 37°C for 4 hours [4].
mRNA Processing: After dephosphorylation, purify mRNA and adjust concentration to 100ng/μl in nuclease-free water. Verify product quality using 1% agarose gel electrophoresis with GelRed staining in 1x TBE buffer [4].
Self-Replicating RNA (srRNA) Synthesis Protocol:
- Plasmid Preparation: Use T7-VEE-OKS-iM plasmids containing coding sequences for Oct4, Sox2, Klf4, cMyc, and non-structural proteins (nsP1-nsP4) of VEE virus. Linearize DNA templates using FastDigest MluI restriction enzyme (36μg plasmid incubated with 5U enzyme for 3h at 37°C) [4].
IVT and Capping: Perform IVT using RiboMAX Large-Scale Production System T7 Kit with 10μg template DNA and 40U RNase Inhibitor. Perform 5′-end capping using ScriptCap Cap1 Capping System followed by 3′-end polyadenylation with A-Plus Poly(A) Polymerase Tailing Kit [4].
Purification and Quality Control: Purify srRNA following each reaction step using ISOLATE II RNA Mini Kit. Analyze length and purity by 1% agarose gel electrophoresis with 2.2M formaldehyde in 1x MOPS buffer at 100V for 60 minutes [4].
Cell Transfection and Reprogramming
Cell Culture Preparation:
- Culture neonatal human foreskin fibroblasts (NuFFs) in DMEM high glucose supplemented with 10% FBS, 1x GlutaMAX, 10mM HEPES, and 50μg/ml gentamicin [4].
- Prepare inactivated feeder cells by treating NuFFs or mouse embryonic fibroblasts with 10mg/ml mitomycin C, then plate on 0.1% gelatin-coated wells [4].
Transient Reprogramming Protocol:
- Mild Trypsinization: For synthetic mRNA approach, detach cells at approximately 70% confluency using 0.04% trypsin/0.03% EDTA, then add trypsin neutralizing solution [4].
mRNA Transfection: Transferd cells daily with 1.2μg synthetic mRNAs or with a single transfection of 1μg srRNA. Include interferon inhibitor B18R in the reprogramming medium to prevent innate immune response and cytotoxicity [4].
Reprogramming Schedule: For partial reprogramming, transferd cells with OSKMLN mRNAs for 4 consecutive days, then analyze 2 days after interruption [12]. For complete iPSC generation, continue daily transfections for 12-15 days [12].
Monitoring: For srRNA containing GFP encoding sequence, monitor reprogramming progress and transfection efficiency through GFP expression [4].
Signaling Pathways and Molecular Dynamics
The Scientist's Toolkit: Essential Research Reagents
Table 4: Essential Research Reagents for Transient mRNA Reprogramming
| Reagent Category | Specific Product | Function in Reprogramming |
|---|---|---|
| Reprogramming Factors | Synthetic modified mRNA (OSKMLN: OCT4, SOX2, KLF4, c-MYC, LIN28, NANOG) | Induction of pluripotency; epigenetic resetting |
| Immune Suppression | B18R protein (interferon inhibitor) | Prevents innate immune response to transfected mRNA |
| Cell Culture Supplement | Modified nucleosides (Pseudouridine, 5-methylcytidine) | Enhances mRNA stability; reduces immune recognition |
| Delivery System | Lipid nanoparticles or electroporation | Efficient intracellular mRNA delivery |
| Quality Control | Agarose gel electrophoresis with GelRed staining | Verifies mRNA integrity and purity |
| Plasmids | T7-VEE-OKS-iM plasmid (for srRNA) | Template for self-replicating RNA production |
| Enzymes for IVT | RiboMAX Large-Scale Production System T7 Kit | High-yield in vitro transcription |
| Capping System | ScriptCap Cap1 Capping System | Adds 5' cap structure for improved translation |
| Polyadenylation Kit | A-Plus Poly(A) Polymerase Tailing Kit | Adds 3' poly(A) tail for mRNA stability |
Transient mRNA expression represents a transformative approach for resetting epigenetic and transcriptional networks without genomic integration. The technology leverages precisely controlled expression of reprogramming factors to reverse age-associated epigenetic marks, restore youthful transcriptional profiles, and ameliorate multiple hallmarks of cellular aging while maintaining cellular identity. The molecular mechanisms involve staged epigenetic remodeling, beginning with rapid changes to chromatin accessibility and DNA methylation patterns, followed by establishment of youthful transcriptional networks. As research advances, optimizing mRNA delivery systems, enhancing translation efficiency, and refining transient expression protocols will further establish this technology as a cornerstone of regenerative medicine and aging research. The non-integrative nature, precision, and safety profile of transient mRNA reprogramming position it as an invaluable tool for both basic research and therapeutic development in pluripotency and cellular rejuvenation.
The advent of induced pluripotent stem cells (iPSCs) has revolutionized regenerative medicine, disease modeling, and drug discovery. A critical advancement in this field has been the development of non-integrative delivery systems that overcome the significant safety concerns associated with early viral vector approaches, which posed risks of insertional mutagenesis and tumorigenesis. Among the most prominent of these advanced systems are mRNA transfection, Sendai virus (SeV) vectors, and episomal plasmids. Each of these technologies enables the transient expression of reprogramming factors without genomic integration, yet they differ substantially in their mechanisms, efficiency, and practical application. This whitepaper provides an in-depth technical analysis of these three key delivery systems, framing their use within the broader context of non-integrative mRNA technology for pluripotency research. It is designed to equip researchers and drug development professionals with the quantitative data, procedural protocols, and strategic insights necessary to select and implement the optimal system for their specific experimental or therapeutic goals.
Technical Comparison of Delivery Systems
The following tables provide a consolidated summary of the performance characteristics and key attributes of the three non-integrative delivery systems, synthesizing data from comparative studies and recent applications.
Table 1: Performance Metrics of Non-Integrative Reprogramming Methods [15]
| Performance Metric | mRNA Transfection | Sendai Virus (SeV) | Episomal Plasmids |
|---|---|---|---|
| Reprogramming Efficiency | 2.1% (Highest) | 0.077% | 0.013% (Lowest) |
| Experimental Success Rate | 27% (Improves to 73-100% with miRNA booster) | 94% | 93% |
| Typical Hands-on Time | ~8 hours | ~3.5 hours | ~4 hours |
| Time to Colony Picking | ~14 days | ~26 days | ~20 days |
| Aneuploidy Rate | 2.3% (Lowest) | 4.6% | 11.5% |
| Transgene Persistence | Short-lived (days) | Lost by passage 9-11 in most lines | Retained in ~33% of lines at passage 9-11 |
Table 2: Key Characteristics and Research Applications [15] [16] [17]
| Characteristic | mRNA Transfection | Sendai Virus (SeV) | Episomal Plasmids |
|---|---|---|---|
| Mechanism of Action | Daily transfection of in vitro transcribed mRNAs encoding factors | Cytoplasmic, non-integrating RNA virus; transduces target cells | Epstein-Barr virus-derived plasmids replicating episomally |
| Key Reprogramming Factors | OSKM + LIN28A + GFP | OSKM | OCT4, SOX2, KLF4, LMYC, LIN28A + shP53 |
| Genomic Integration | None | None; exclusively cytoplasmic replication | Low-rate integration possible; primarily extrachromosomal |
| Advantages | No risk of integration; fastest kinetics; high efficiency | Broad cell tropism; high transduction efficiency; reliable | Simple delivery (transfection); no viral components |
| Disadvantages | High workload; massive cell death in some samples; requires immune suppression | Requires screening for viral clearance; longer timeline | Lower efficiency; potential for plasmid retention |
| Ideal Application | Clinical-grade iPSCs where speed and integration-free status are critical | Robust reprogramming of difficult-to-transfect cells; basic research | Studies avoiding viral vectors; facilities with standard transfection expertise |
Detailed Methodologies and Experimental Protocols
mRNA Transfection Protocol
The mRNA transfection method involves the daily delivery of synthetic mRNAs encoding reprogramming factors into somatic cells, triggering their reprogramming into iPSCs.
- Key Reagents: mRNA reprogramming kit (e.g., from Stemgent); immune suppressants (e.g., B18R interferon inhibitor); transfection reagent; fibroblast culture medium.
Procedure:
- Day 0: Seed Somatic Cells: Plate human fibroblasts (e.g., neonatal BJ or patient-derived PS lines) at a density of 5 x 10^4 cells per well in a 6-well plate.
- Days 1-?: Daily Transfection:
- Prepare mRNA-lipid complexes per manufacturer's instructions. A typical cocktail includes mRNAs for OCT4, SOX2, KLF4, cMYC, LIN28A, and a GFP reporter.
- Replace cell culture medium with fresh medium containing an immune suppressor.
- Add the mRNA-lipid complexes to the cells. Transfection efficiency can be monitored via GFP expression.
- Repeat this process daily until colony formation is observed (typically ~14 days).
- Colony Picking and Expansion: Once compact, ESC-like colonies appear, manually pick and transfer them to feeder-coated plates for expansion under standard hiPSC culture conditions.
Critical Considerations: This protocol is labor-intensive and can trigger innate immune responses, leading to significant cell death in some samples. The use of a microRNA (miRNA) Booster Kit can significantly improve the success rate from 27% to 73-100% for refractory samples [15].
Sendai Virus (SeV) Transduction Protocol
The Sendai virus is an RNA virus that replicates in the cytoplasm without integrating into the host genome, making it a safe and efficient vector for reprogramming.
- Key Reagents: SeV-based vector kit (e.g., Cytotune from Life Technologies); appropriate cell culture medium.
Procedure:
- Day 0: Seed Target Cells: Plate the somatic cells (e.g., fibroblasts) at an appropriate density (e.g., 1 x 10^5 cells per well in a 6-well plate).
- Day 1: Viral Transduction:
- Thaw the SeV particles (encoding OCT4, SOX2, KLF4, cMYC) quickly and keep on ice.
- Replace the cell medium with a minimal volume of medium containing the appropriate multiplicity of infection (MOI) of each viral vector. A typical approach is a cocktail of separate vectors for each factor.
- Incubate the cells for several hours (e.g., 4-6 hours) before adding more complete medium.
- Days 2-26: Culture and Monitor:
- Change the medium regularly.
- Monitor for the appearance of reprogrammed colonies, which typically emerge around day 26.
- The virus is naturally diluted and lost through cell passaging. Screening for virus clearance via RT-PCR is recommended by passages 9-11 [15].
- Colony Picking: Pick and expand virus-free colonies.
Note on SeVdp Vectors: Recent advances use replication-defective, persistent SeVdp vectors. These vectors offer robust, high-level transgene expression with minimal cytopathic effects and are particularly useful for direct reprogramming, as demonstrated in the induction of chondrocytes from fibroblasts [18].
Episomal Plasmid Transfection Protocol
Episomal plasmids utilize elements from the Epstein-Barr virus to replicate extrachromosomally in dividing cells, providing transient transgene expression.
- Key Reagents: Episomal plasmids (e.g., encoding OCT4, SOX2, KLF4, LMYC, LIN28A, and shP53); transfection reagent (e.g., Lipofectamine); nucleofector device and kit (for electroporation).
- Procedure:
- Day 0: Prepare Cells: Harvest and count somatic cells. For electroporation, use 1-2 x 10^6 cells per transfection.
- Day 1: Plasmid Delivery:
- Electroporation (Recommended): Co-electroporate a combination of episomal plasmids (e.g., 2-3 different plasmids carrying the reprogramming factors) into the cells using a nucleofection system optimized for the cell type.
- Lipofection: Alternatively, form DNA-lipid complexes and add them to the cells.
- Days 2-20: Culture and Passage:
- 24 hours post-transfection, transfer the cells onto feeder layers.
- Culture the cells, passaging as needed. Colonies typically appear and are ready for picking around day 20.
- Colony Screening: Expand picked colonies and screen for the loss of the EBNA1 plasmid sequence via PCR. At passage 10, roughly one-third of lines may retain the plasmid and should be excluded from further use [15].
Visualizing the Experimental Workflows
The following diagrams illustrate the core workflows and mechanisms for each delivery system, providing a logical map for experimental planning.
mRNA Transfection Workflow
Sendai Virus (SeV) Transduction Workflow
Episomal Plasmid Transfection Workflow
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Non-Integrative Reprogramming [15] [18]
| Reagent / Kit Name | Function | Application / Note |
|---|---|---|
| Stemgent mRNA Reprogramming Kit | Provides synthetic mRNAs for reprogramming factors (OSKM, LIN28) | Core reagent for mRNA transfection protocol; requires daily transfection |
| miRNA Booster Kit (Stemgent) | Improves reprogramming efficiency and success rate | Used in combination with mRNA kit to overcome cell death in refractory samples |
| Cytotune iPS Sendai Reprogramming Kit (Life Technologies) | Provides SeV particles for the four Yamanaka factors (OSKM) | Kit-based solution for SeV reprogramming; includes separate amps for each factor |
| SeVdp (Delta F) Vectors | Replication-defective, persistent Sendai virus vectors | Minimizes cytopathic effects; allows for stable, high-level transgene expression |
| Episomal Plasmids (e.g., pCEP4-based vectors) | DNA vectors for reprogramming factor expression | Typically used in sets of 2-3 plasmids; often include LMYC and shP53 for higher efficiency |
| H2B-mKO2 Tagged Reporter Plasmids | Fluorescent reporter to identify plasmid-retaining colonies | Enables visual screening for episomal plasmid loss during iPSC expansion |
The strategic selection of a non-integrative delivery system is a cornerstone of successful iPSC generation. mRNA transfection, Sendai virus vectors, and episomal plasmids each present a distinct profile of advantages and limitations, making them suitable for different research contexts. The choice hinges on the specific priorities of the project: mRNA transfection offers the highest speed and a clean integration-free profile but demands significant hands-on effort; Sendai virus provides robust efficiency and reliability but requires monitoring for viral clearance; and episomal plasmids offer a simple, viral-free workflow but at lower efficiency and with a need for plasmid clearance checks. As the field advances, technologies like deep learning-optimized mRNA codons and refined centromeric plasmids promise to further enhance the efficiency and safety of these systems [16] [19] [20]. By leveraging the quantitative data, detailed protocols, and strategic insights contained in this whitepaper, researchers can effectively harness these powerful technologies to drive innovation in pluripotency research and therapeutic development.
The advent of induced pluripotent stem cell (iPSC) technology, pioneered by Takahashi and Yamanaka's seminal work, demonstrated that somatic cells could be reprogrammed into a pluripotent state using defined transcription factors [21]. The initial reprogramming methodologies relied heavily on integrating viral vectors, such as retroviruses and lentiviruses, to deliver the essential reprogramming factors (OCT4, SOX2, KLF4, and c-MYC) [21]. While effective, a significant safety concern inherent to these methods is insertional mutagenesis—the random integration of viral DNA into the host genome which can disrupt tumor suppressor genes, activate oncogenes, or cause other genomic alterations that increase the risk of tumorigenesis in derived cells [21] [22]. This risk presents a major barrier to the clinical translation of iPSC-based therapies.
Non-integrative mRNA technology has emerged as a powerful alternative, completely eliminating the risk of insertional mutagenesis by delivering reprogramming factors as transient messenger RNA molecules that do not enter the nucleus or interact with the host genome [16]. This technical guide examines the core mechanisms, advantages, and methodological protocols of non-integrating mRNA reprogramming, positioning it as a cornerstone for the development of clinically safe pluripotent stem cells.
Technical Mechanisms: How Non-Integrating mRNA Methods Ensure Genomic Safety
Non-integrating mRNA reprogramming leverages synthetic, modified messenger RNA to transiently express reprogramming factors in somatic cells. The core mechanism hinges on the natural function of mRNA: once delivered to the cell cytoplasm, it is directly translated into protein by the host ribosomes without any nuclear entry or interaction with chromosomal DNA [16] [23]. This fundamental difference from DNA-based methods is the basis of its enhanced safety profile.
- Cytoplasmic Translation and Nuclear Function: The translated reprogramming proteins (OCT4, SOX2, KLF4, c-MYC) translocate to the nucleus where they perform their function of activating pluripotency networks. The mRNA itself remains in the cytoplasm and is eventually degraded by natural cellular processes [16] [24].
- Transient Expression Kinetics: Unlike integrated proviruses, which can lead to sustained and uncontrolled expression of reprogramming factors, mRNA expression is transient, typically lasting only a few days. This necessitates repeated transfections but prevents lingering expression of oncogenes like c-MYC, which is known to promote tumor formation if persistently expressed [16] [21].
- Avoidance of Genome-Wide Disruption: By foregoing genomic integration, mRNA methods eliminate the risks of insertional mutagenesis, such as the disruption of coding regions or tumor suppressor genes, and the potential for sustained DNA damage responses that can be triggered by the integration events themselves [25] [24].
Table 1: Key Safety and Efficiency Metrics of Non-Integrative Reprogramming Methods
| Method | Genomic Integration Risk | Typical Reprogramming Efficiency | Key Safety Features | Primary Clinical Applicability |
|---|---|---|---|---|
| mRNA Transfection | None [16] | Moderate to High [16] | No integration, controlled kinetics, no anti-vector immunity [16] [21] | Clinical-grade iPSC generation [16] |
| Sendai Virus (SeV) | None (cytoplasmic RNA virus) [16] [21] | High [16] | Non-integrating, eventually diluted by cell division [21] | GMP-compliant iPSC generation [16] |
| Episomal Plasmids | Very Low (non-integrating, but theoretical risk) [22] | Low to Moderate [22] | Non-viral, plasmid is typically lost over cell divisions [22] | Research and preclinical development [22] |
| Integrating Retrovirus | High (random integration) [21] [22] | High | N/A (Obsolete for clinical use due to safety profile) | Foundational research only |
Established Protocols for mRNA-Based Reprogramming
This section provides a detailed, actionable protocol for generating iPSCs using synthetic mRNA, based on established and optimized procedures.
Protocol: iPSC Generation via Daily mRNA Transfection
Objective: To reprogram human somatic cells (e.g., dermal fibroblasts or peripheral blood mononuclear cells) into induced pluripotent stem cells using repeated transfections of synthetic mRNA encoding reprogramming factors.
Materials and Reagents:
- Somatic Cells: Human neonatal or adult dermal fibroblasts.
- mRNA Cocktail: A commercially available kit or custom-synthesized, modified mRNA molecules encoding human OCT4, SOX2, KLF4, c-MYC, and LIN28. These mRNAs feature nucleotide modifications (e.g., pseudouridine) to reduce innate immune recognition and enhance stability [16] [23].
- Transfection Reagent: A specialized transfection reagent designed for efficient mRNA delivery.
- Cell Culture Media: Fibroblast growth medium, iPSC reprogramming medium, and essential iPSC maintenance medium (e.g., mTeSR or E8).
- Matrix: Recombinant human vitronectin or Matrigel for coating culture vessels.
Methodology:
- Day 0: Cell Plating: Plate human fibroblasts at a defined density (e.g., 20,000 cells per well of a 12-well plate) in fibroblast growth medium. Cells should be attached and have a confluence of approximately 50-70% at the time of the first transfection.
- Days 1-7: Daily mRNA Transfection:
- Prepare the mRNA-lipid complex by combining the mRNA cocktail with the transfection reagent in a serum-free medium, following the manufacturer's optimized ratio.
- Incubate the mixture for 10-15 minutes at room temperature to allow complex formation.
- Aspirate the culture medium from the cells and wash once with PBS.
- Add the mRNA-lipid complex dropwise to the cells in fresh, serum-free reprogramming medium.
- Incubate cells for 4-6 hours, after which replace the transfection medium with fresh reprogramming medium supplemented with B18R interferon inhibitor (to mitigate immune responses against double-stranded RNA contaminants) [16].
- Days 7-21: Colony Observation and Picking:
- Around day 7, change the medium to a defined iPSC maintenance medium.
- Between days 14-21, distinct, compact iPSC colonies with defined borders and high nucleus-to-cytoplasm ratio will emerge.
- Manually pick individual colonies using a sterile pipette tip or use enzymatic methods to dissect and transfer colonies onto fresh matrix-coated plates for expansion.
Critical Steps and Troubleshooting:
- Cell Density: Optimal cell density at the first transfection is critical for successful reprogramming efficiency.
- mRNA Quality and Purity: Ensure mRNA is of high purity and integrity to minimize activation of the innate immune system (e.g., PKR and OAS pathways), which can halt translation and induce apoptosis [16] [23].
- Interferon Suppression: The use of B18R protein is crucial for enhancing cell survival and reprogramming efficiency by counteracting the innate immune response.
Diagram 1: mRNA Reprogramming Workflow
Comparative Analysis: Signaling and Immune Pathway Activation
Non-integrating mRNA methods interact with cellular machinery in a fundamentally different way compared to viral methods, particularly in their engagement with innate immune and DNA damage response pathways.
Diagram 2: Safety Pathway Comparison
- Innate Immune Recognition: Exogenous mRNA can be recognized by cytoplasmic pattern recognition receptors (e.g., RIG-I) and endosomal Toll-like receptors (TLRs), triggering a type I interferon response [25] [23]. This response, while a potential hurdle, can be effectively managed. The use of nucleoside-modified mRNA (e.g., pseudouridine) reduces receptor binding, and the supplementation of interferon inhibitors like B18R during reprogramming ensures high cell viability and protein expression [16] [23].
- Absence of DNA Damage Response: Viral vector integration often causes double-strand breaks (DSBs) in the host DNA, activating the p53-mediated DNA damage response pathway. This can lead to cell senescence or apoptosis, thereby limiting reprogramming efficiency, or conversely, selecting for cells with compromised p53 function, which carries oncogenic risk [21]. mRNA methods completely bypass this activation, as no DSBs are incurred, supporting a safer and more controlled reprogramming process [16].
Table 2: The Scientist's Toolkit: Essential Reagents for mRNA Reprogramming
| Reagent / Solution | Function and Mechanism | Key Characteristic |
|---|---|---|
| Nucleoside-Modified mRNA | Serves as the transient template for reprogramming factor protein synthesis. Modified bases (e.g., pseudouridine) prevent innate immune recognition [16] [23]. | High translational efficiency, reduced immunogenicity. |
| Lipid-Based Transfection Reagent | Encapsulates and delivers mRNA across the cell membrane via endocytosis and endosomal escape [16]. | High efficiency, low cytotoxicity formulations. |
| B18R Interferon Inhibitor | A recombinant protein that binds and neutralizes type I interferons, blocking the antiviral state and enhancing cell survival during repeated transfections [16]. | Critical for multi-day transfection protocols. |
| Vitronectin-coated Plates | Provides a defined, xeno-free extracellular matrix substrate for the attachment and growth of emerging iPSC colonies [21]. | Defined, GMP-compatible substrate. |
| Small Molecule Cocktails (e.g., A-83-01) | Enhances reprogramming efficiency by inhibiting TGF-β signaling and other pathways that maintain somatic cell identity [16]. | Synergistic with mRNA, improves kinetics. |
The adoption of non-integrating mRNA technology represents a paradigm shift in pluripotency research, directly addressing the critical safety concern of insertional mutagenesis that has long hindered the clinical translation of iPSCs. By enabling transient, high-efficiency expression of reprogramming factors without genomic alteration, this method facilitates the generation of footprint-free iPSCs that are biologically closer to a pristine embryonic state. As the field advances, the convergence of mRNA technology with other innovations—such as CRISPR-Cas9 gene editing (where mRNA is also used to deliver Cas9 protein) and AI-guided differentiation protocols—is poised to further accelerate the development of safe and effective personalized cell therapies, regenerative medicine applications, and high-fidelity disease models [16] [21]. The continued refinement of mRNA design, delivery systems, and manufacturing protocols under GMP standards will solidify its role as a foundational technology for the next generation of clinical-grade pluripotent stem cells.
Protocols and Implementations: Applying mRNA Technology for iPSC Generation and Differentiation
The discovery that somatic cells can be reprogrammed into induced pluripotent stem cells (iPSCs) using defined factors has revolutionized regenerative medicine and disease modeling [26]. Traditional methods relying on viral vectors for factor delivery pose significant clinical risks due to genomic integration and potential insertional mutagenesis [21]. mRNA-based technology has emerged as a superior non-integrative approach that combines high reprogramming efficiency with enhanced safety profiles [16] [27]. This transient delivery method avoids permanent genetic alterations, making it particularly valuable for clinical applications and pharmaceutical development [9] [28].
Unlike early reprogramming methods that used retroviral or lentiviral vectors, mRNA reprogramming delivers synthetic mRNAs encoding key transcription factors to somatic cells without integrating into the host genome [21] [26]. The technology leverages modified nucleobases in the mRNA construct to reduce innate immune recognition while maintaining high protein expression levels [27]. This whitepaper provides a comprehensive technical guide to implementing mRNA-based somatic cell reprogramming, with detailed protocols, optimization parameters, and quality control measures essential for successful iPSC generation.
Core Principles and Advantages
Fundamental Mechanisms
The mRNA reprogramming process involves the introduction of in vitro transcribed mRNAs encoding the core pluripotency factors OCT4, SOX2, KLF4, and c-MYC (OSKM) into somatic cells [26] [27]. These mRNAs are translated into proteins that initiate a cascade of transcriptional and epigenetic changes, ultimately driving the cells toward a pluripotent state. The process typically requires repeated transfections over several days to maintain sufficient levels of reprogramming factors as the cells undergo this identity transformation [27].
The mechanism relies on the cell's native translational machinery to produce the reprogramming proteins, avoiding the unpredictability of viral integration sites and transgene silencing issues associated with DNA-based methods [21]. The non-integrating nature of this technology ensures that the reprogrammed iPSCs are "footprint-free," meaning they carry no foreign genetic material, which is crucial for clinical applications [28].
Key Advantages Over Alternative Methods
Table 1: Comparison of mRNA Reprogramming with Other Methods
| Parameter | mRNA Method | Viral Methods | Episomal Plasmid | Sendai Virus |
|---|---|---|---|---|
| Genomic Integration | None | High | Low | None |
| Reprogramming Efficiency | High (up to 90.7%) [27] | Moderate | Low | Moderate to High |
| Reprogramming Time | 2-4 weeks | 3-4 weeks | 4-6 weeks | 3-4 weeks |
| Safety Profile | Excellent | Poor (tumor risk) | Good | Good |
| Clinical Applicability | High | Low | Moderate | High |
| Technical Difficulty | High | Moderate | Low | Moderate |
The mRNA platform provides precision, safety, and transience in directing cellular behavior [9]. Its non-integrative nature and controllable strategy for expressing therapeutic proteins make it particularly suitable for clinical translation [9] [16]. Modern reprogramming methods have significantly reduced genomic alterations through these safer non-integrative approaches, replacing traditional viral methods for generating clinical-grade iPSCs [16].
Technical Workflow
Pre-Reprogramming Preparation
Starting Cell Culture
- Cell Type Selection: Human primary fibroblasts from neonatal foreskin or adult skin biopsies are commonly used [27]. Ensure cells are at low passage (passage 3-6) and have >95% viability before reprogramming.
- Culture Conditions: Maintain fibroblasts in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 1% GlutaMAX, and 1% non-essential amino acids [27].
- Quality Control: Confirm cell identity through morphology and biomarker expression. Test for mycoplasma contamination.
mRNA Preparation
- mRNA Constructs: Use synthetic mRNAs encoding OCT4, SOX2, KLF4, c-MYC, LIN28, and NANOG [27]. Some protocols use a modified OCT4 (M3O) with the MyoD transactivation domain for enhanced efficiency [27].
- Nucleoside Modifications: Incorporate modified nucleobases (e.g., pseudouridine, 5-methylcytidine) to reduce innate immune recognition [27].
- miRNA Enhancement: Supplement with ESC-specific miRNA-367/302s as mature miRNA mimics to synergistically enhance reprogramming efficiency [27].
Optimized Reprogramming Protocol
The following workflow has been optimized for high-efficiency reprogramming of human primary fibroblasts:
Diagram 1: mRNA Reprogramming Workflow
Detailed Step-by-Step Procedure
Day 0: Cell Seeding
- Harvest fibroblasts using standard trypsinization procedure.
- Count cells using automated counter or hemocytometer.
- Plate 500 cells per well of a 6-well plate in fibroblast medium.
- Incubate overnight at 37°C, 5% CO₂.
Day 1: First Transfection
- Prepare transfection complex A: Dilute 600 ng of 5fM3O mod-mRNA cocktail and 20 pmol of miRNA-367/302s mimics in 125 μL of Opti-MEM adjusted to pH 8.2.
- Prepare transfection complex B: Dilute 6 μL of Lipofectamine RNAiMAX in 125 μL of Opti-MEM pH 8.2.
- Incubate both complexes for 5-10 minutes at room temperature.
- Combine complexes A and B, mix gently, and incubate for 20-30 minutes.
- Add complex mixture dropwise to cells in 2 mL of KOSR reprogramming medium.
- Incubate cells at 37°C, 5% CO₂ for 24 hours.
Days 3, 5, 7, 9, 11, 13: Repeated Transfections
- Repeat the transfection procedure every 48 hours for a total of 7 transfections.
- Change medium 4-6 hours after each transfection to reduce cytotoxicity.
- Monitor cell morphology daily for emergence of compact, ESC-like colonies.
Days 7-21: Colony Monitoring and Picking
- Beginning around day 7, monitor for emergence of TRA-1-60-positive colonies.
- Between days 14-21, pick individual colonies using sterile pipette tips.
- Transfer colonies to matrigel-coated plates with mTeSR or Essential 8 medium.
- Expand colonies for characterization and banking.
Critical Optimization Parameters
Table 2: Key Optimization Parameters for mRNA Reprogramming
| Parameter | Optimal Condition | Effect on Reprogramming | Reference |
|---|---|---|---|
| Cell Seeding Density | 500 cells/well (6-well plate) | Prevents contact inhibition, allows more cell cycles | [27] |
| Transfection Interval | Every 48 hours | Maintains consistent factor expression | [27] |
| Transfection Buffer pH | Opti-MEM pH 8.2 | Increases transfection efficiency to ~65% | [27] |
| mRNA Dose | 600 ng 5fM3O + 20 pmol miRNAs | Balances expression and cytotoxicity | [27] |
| miRNA Supplementation | miRNA-367/302s mimics | Synergistic enhancement, 90.7% efficiency | [27] |
| Minimum Transfections | 3 sessions | Essential for complete reprogramming | [27] |
The Scientist's Toolkit: Essential Reagents and Materials
Table 3: Key Research Reagent Solutions for mRNA Reprogramming
| Reagent Category | Specific Product/Component | Function in Reprogramming |
|---|---|---|
| Reprogramming mRNAs | 5fM3O mod-mRNA cocktail (OCT4, SOX2, KLF4, c-MYC, LIN28, NANOG) | Core factors inducing pluripotency |
| Enhancing miRNAs | miRNA-367/302s mimics | Synergistically improves efficiency & colony formation |
| Transfection Reagent | Lipofectamine RNAiMAX | Efficient RNA delivery with low cytotoxicity |
| Transfection Buffer | Opti-MEM pH 8.2 | Optimized buffer for high transfection efficiency |
| Reprogramming Medium | KnockOut Serum Replacement (KOSR) Medium | Supports reprogramming while maintaining cell viability |
| Culture Matrix | Matrigel or Recombinant Laminin-521 | Provides substrate for iPSC colony attachment & growth |
| iPSC Maintenance | mTeSR1 or Essential 8 Medium | Defined medium for pluripotent stem cell culture |
Signaling Pathways and Molecular Mechanisms
The reprogramming process involves complex signaling pathways that are activated by the introduced transcription factors:
Diagram 2: Key Signaling Pathways in Reprogramming
Key Molecular Events
- Transcriptional Activation: OCT4 and SOX2 directly activate endogenous pluripotency genes including NANOG, while suppressing somatic cell programs [1] [26].
- Epigenetic Remodeling: KLF4 recruits chromatin modifiers that facilitate the opening of repressed pluripotency loci through DNA demethylation and histone modifications [1].
- Metabolic Reprogramming: c-MYC drives a shift from oxidative phosphorylation to glycolysis, characteristic of pluripotent cells [1] [26].
- Signaling Pathway Modulation: The process involves activation of Wnt and suppression of BMP signaling, mimicking the native embryonic stem cell niche [16].
Quality Control and Characterization
Essential Validation Assays
- Pluripotency Marker Expression: Confirm expression of TRA-1-60, TRA-1-81, SSEA4 through immunocytochemistry and flow cytometry [27].
- Gene Expression Analysis: Verify upregulation of endogenous pluripotency genes (OCT4, SOX2, NANOG) via RT-qPCR.
- Trilineage Differentiation Potential: Demonstrate differentiation into ectoderm, mesoderm, and endoderm lineages through in vitro differentiation and teratoma formation assays [28].
- Karyotype Analysis: Perform G-banding karyotyping to ensure genetic normality (46 chromosomes).
- Identity Verification: Confirm match to donor somatic cells through STR profiling.
Troubleshooting Common Issues
- Low Efficiency: Optimize transfection buffer pH, ensure high-quality mRNA, and confirm appropriate cell seeding density.
- High Cell Death: Reduce mRNA dose, increase medium changes post-transfection, and verify cell viability before starting.
- No Colony Formation: Check reprogramming mRNA activity, confirm appropriate culture conditions, and validate primary cell quality.
- Spontaneous Differentiation: Improve picking technique, optimize passage timing, and ensure quality of extracellular matrix coating.
Applications in Research and Therapy
The mRNA-reprogrammed iPSCs have broad applications across multiple domains:
- Disease Modeling: Patient-specific iPSCs enable in vitro modeling of neurodegenerative diseases, cardiac disorders, and genetic conditions [1] [29].
- Drug Discovery and Screening: iPSC-derived cells provide human-relevant platforms for compound testing and safety pharmacology [29] [21].
- Cell Replacement Therapy: Clinical trials are underway using iPSC-derived cells for Parkinson's disease, age-related macular degeneration, and cardiac repair [21].
- Cancer Immunotherapy: iPSC-derived natural killer (NK) cells and chimeric antigen receptor (CAR)-T cells offer off-the-shelf cancer treatments [1] [29].
mRNA-based somatic cell reprogramming represents a robust, efficient, and clinically relevant method for generating integration-free iPSCs. The protocol outlined in this whitepaper, with its optimized transfection conditions, miRNA supplementation, and culture parameters, enables researchers to achieve reprogramming efficiencies exceeding 90% while maintaining the genetic integrity essential for downstream applications. As non-integrative mRNA technology continues to advance, it promises to accelerate the translation of iPSC-based therapies from research laboratories to clinical practice, ultimately enabling personalized regenerative medicine approaches for a wide range of degenerative diseases.
The generation of clinical-grade induced pluripotent stem cells (iPSCs) represents a cornerstone in the advancement of regenerative medicine and cell-based therapies. Unlike research-grade iPSCs, clinical-grade lines must adhere to rigorous Good Manufacturing Practice (GMP) standards and quality control measures to ensure their safety, efficacy, and consistency for human therapeutic applications [30] [31]. These cells are characterized by their derivation under fully defined, xeno-free conditions using integration-free reprogramming methods, with comprehensive documentation and rigorous safety testing throughout the manufacturing process [31] [32]. The transition toward clinical-grade iPSCs has been significantly accelerated by the development of non-integrative mRNA reprogramming technology, which offers a precise, footprint-free method for inducing pluripotency without genomic modification, thereby addressing critical safety concerns associated with earlier viral methods [9] [28] [16].
The fundamental distinction between clinical and research-grade iPSCs lies in the comprehensive regulatory framework governing their production. According to international consensus workshops, clinical-grade lines require agreement on critical quality attributes and standardized assays to demonstrate comparability across lines derived from different individuals and facilities [30]. This includes strict adherence to GMP principles throughout the entire process—from donor screening and tissue acquisition to reprogramming, characterization, and banking [31] [32]. The emergence of non-integrative mRNA technology has been particularly transformative for clinical applications, as it eliminates the risk of insertional mutagenesis while providing a controlled, reproducible reprogramming process compatible with regulatory requirements for clinical use [28] [33] [16].
Fundamental Principles of GMP Compliance
Core GMP Requirements for iPSC Manufacturing
Good Manufacturing Practice establishes a comprehensive framework to ensure the quality, safety, and consistency of iPSC lines intended for clinical applications. The core principles encompass several critical aspects of production. Documentation and traceability require that all materials, procedures, and personnel involved in manufacturing are meticulously documented to ensure full traceability from donor source to final cell bank [31] [32]. This includes maintaining detailed batch records, standard operating procedures, and chain of custody documentation. Facility and environmental controls mandate that all manufacturing processes occur in controlled environments with appropriate air quality, monitoring, and cleanliness standards to prevent contamination [31]. Personnel training and qualification ensure that all staff are thoroughly trained in GMP principles and specific technical procedures, with training records maintained and regularly reviewed [32].
A cornerstone of GMP compliance is the implementation of a Quality Management System that encompasses all aspects of production, including quality control testing, deviation management, change control, and release specifications [30] [32]. Additionally, material control and qualification requires that all raw materials, reagents, and components are properly qualified, stored, and tracked according to established protocols, with particular emphasis on using xeno-free, clinically-approved materials [31] [32]. The equipment validation and maintenance principle dictates that all equipment used in manufacturing must be properly validated, calibrated, and maintained according to predefined schedules to ensure consistent performance [32]. Finally, lot-to-lot consistency and specification establishes that each manufactured lot must meet predefined release specifications and demonstrate consistency with previous lots [30].
Donor Eligibility and Starting Material Considerations
The selection of appropriate donor material represents the first critical step in generating clinical-grade iPSCs. Donors must undergo comprehensive screening according to national and international "Tissue Donor Guidance" regulations, which typically includes medical history review and infectious disease testing [31]. Written informed consent specifically covering clinical and commercial use of derived cells is essential, with no financial benefits involved in the donation process [31]. Umbilical cord-derived mesenchymal stromal cells have emerged as an ideal starting material due to their immature characteristics, limited environmental exposure, and availability from GMP-compliant perinatal tissue banks [32]. These cells offer advantages including known family and medical histories of donors, reduced time and costs associated with personalized treatments, and GMP-compliant sourcing [32].
Alternative donor sources such as clinical-grade human foreskin fibroblasts have also been successfully utilized, with isolation and culture performed using xeno-free reagents in GMP-grade laboratories [31]. These parental cells must be confirmed negative for mycoplasma and specific pathogenic microorganisms, with biological safety validated by national control agencies [31]. The use of well-characterized starting materials from eligible donors provides a critical foundation for generating safe, clinically applicable iPSC lines that meet regulatory requirements across multiple jurisdictions, including the US FDA, European EMA, and Japanese PMDA [33] [32].
Non-Integrative mRNA Reprogramming Technology
Mechanism and Advantages of mRNA Reprogramming
Non-integrative mRNA reprogramming technology represents a groundbreaking approach for generating clinical-grade iPSCs through transient expression of reprogramming factors without genomic integration. This method utilizes engineered messenger RNA constructs that encode key transcription factors—typically OCT4, SOX2, KLF4, and c-MYC (OSKM)—to reprogram somatic cells into pluripotent stem cells [28] [16]. The fundamental mechanism involves introducing these modified mRNA sequences into target cells, where they are translated into functional proteins that initiate and drive the reprogramming process [9] [28]. Unlike viral methods, mRNA reprogramming leaves no genomic footprint as the mRNA is not retained in the cells and cannot integrate into the host genome, thereby eliminating concerns about insertional mutagenesis and providing a significant safety advantage for clinical applications [28] [33].
The technological advantages of mRNA reprogramming are substantial. The controlled expression of reprogramming factors enables precise regulation of reprogramming kinetics, while the rapid turnover of mRNA allows for dynamic adjustment of factor expression through dosing regimens [9] [16]. Additionally, this approach demonstrates high reprogramming efficiency, often generating genetically stable iPSCs with lower rates of genomic abnormalities compared to other methods [33]. The compatibility with clinical applications is high, as the process uses defined components without viral elements, meeting regulatory requirements for clinical-grade cell production [28] [33]. Furthermore, the avoidance of transgene persistence ensures that no exogenous genetic material remains in the resulting iPSCs, addressing critical safety concerns [28] [16].
Experimental Protocol for mRNA Reprogramming
The implementation of mRNA reprogramming requires meticulous protocol execution. The process begins with the preparation of somatic cells, such as human dermal fibroblasts or umbilical cord mesenchymal stromal cells, which are cultured and expanded under xeno-free conditions until they reach 70-80% confluence [31] [32]. Concurrently, mRNA preparation involves diluting the reprogramming factor mRNAs (OCT4, SOX2, KLF4, c-MYC, and optionally LIN28 or other factors) in an appropriate buffer. Some protocols incorporate modified nucleosides such as pseudouridine to reduce innate immune recognition and enhance translation efficiency [9].
The transfection process is typically performed using a lipid-based transfection reagent compatible with clinical applications. The mRNA-lipid complexes are added to the cells daily for approximately 12-18 days, with medium changes 4-6 hours post-transfection to minimize cellular stress [28]. Critical to this process is the optimization of mRNA ratios, as studies indicate that the specific ratio of SOX2 to OCT4 significantly affects reprogramming efficiency and colony quality [34]. Following transfection, colony emergence typically occurs between days 7-10, with iPSC colony picking performed between days 18-25 based on morphological criteria resembling human embryonic stem cells [31].
The expansion and characterization phase involves transferring picked colonies to GMP-compliant, feeder-free culture systems using defined matrices and xeno-free media for expansion [33] [31]. Throughout the process, quality control monitoring includes regular assessment of cell morphology, growth rates, and pluripotency marker expression [33] [31]. This comprehensive protocol enables the generation of footprint-free iPSCs suitable for clinical applications, with successful implementation demonstrated by commercial providers such as Pluristyx and REPROCELL, who utilize proprietary mRNA technologies to produce clinical-grade iPSC lines [28] [33].
Figure 1: mRNA Reprogramming Workflow for Clinical-Grade iPSCs. This diagram illustrates the sequential steps in non-integrative mRNA reprogramming, highlighting key safety features including xeno-free conditions and footprint-free results.
Essential Quality Control Assays
Mandatory Quality Control Testing
Quality control testing for clinical-grade iPSCs encompasses a comprehensive panel of assays designed to verify identity, purity, potency, and safety. The identity testing includes pluripotency verification through flow cytometry analysis of surface markers (TRA-1-60, TRA-1-81, SSEA-4) and intracellular markers (OCT4, NANOG, SOX2), with specification thresholds typically requiring >90% expression for key markers [33] [31]. Additionally, short tandem repeat profiling confirms donor identity and detects cross-contamination, while pluripotency assessment involves directed differentiation into all three germ layers with evaluation of representative markers: ectoderm (PAX6, SOX1), mesoderm (Brachyury, SMA), and endoderm (SOX17, AFP) [33] [31].
For safety testing, sterility assessments are critical and include bacteriology and fungiology culture (14 days) with a specification of no contamination, mycoplasma testing by PCR and/or culture (minimum 28 days) with no detection allowed, and endotoxin testing with a typical specification of <0.5 EU/mL [31]. Viral safety requires testing for adventitious viruses (in vitro and in vivo assays) with no cytopathic effect allowed, and specific pathogen testing including HIV-1/2, HBV, HCV, and others relevant to donor epidemiology [31]. Genetic stability assessment involves G-band karyotyping to ensure normal chromosomal number and structure without major abnormalities, with some facilities additionally performing next-generation sequencing-based oncogenetic analysis to profile genetic variants in over 400 cancer-related genes [33].
Release Criteria and Specifications
The release of clinical-grade iPSC lines for therapeutic applications requires meeting stringent specification criteria across multiple quality attributes. The following table summarizes the standard release criteria for clinical-grade iPSCs based on current guidelines and practices [30] [33] [31]:
Table 1: Standard Release Criteria for Clinical-Grade iPSCs
| Quality Attribute | Test Method | Release Specification | Frequency |
|---|---|---|---|
| Pluripotency | Flow Cytometry | >90% expression of key pluripotency markers (OCT4, SOX2, SSEA-4) | Every cell bank |
| Trilineage Differentiation | Directed Differentiation | Demonstrated differentiation into all three germ layers with appropriate marker expression | At characterization |
| Karyotype | G-band Karyotyping | Normal chromosomal complement (46, XX or XY) without structural abnormalities | Every cell bank |
| Oncogenetic Mutations | NGS Panel | No high-impact mutations in 400+ cancer-related genes | At characterization |
| Sterility | Microbiology Culture | No bacterial or fungal contamination | Every lot |
| Mycoplasma | PCR and/or Culture | Negative | Every cell bank |
| Endotoxin | LAL Test | <0.5 EU/mL | Every lot |
| Viral Safety | PCR/In vitro Assays | Negative for specified adventitious viruses | Donor screening and cell bank |
These release criteria ensure that clinical-grade iPSCs meet the necessary quality standards for use in human therapies. The combination of multiple complementary techniques provides a comprehensive safety assessment, with particular emphasis on genetic integrity through both low-resolution karyotyping and higher-resolution molecular analysis [33]. Additionally, the functional assessment of pluripotency through trilineage differentiation confirms the biological potency of the cells, which is essential for their intended therapeutic applications [33] [31].
Signaling Pathways in Pluripotency and Reprogramming
Molecular Regulation of Pluripotency
The establishment and maintenance of pluripotency in iPSCs are governed by complex signaling pathways that coordinate to regulate the expression of core transcription factors and epigenetic modifiers. The core pluripotency network centers on the transcription factors OCT4, SOX2, and NANOG, which form an interconnected autoregulatory loop that activates genes essential for maintaining pluripotency while suppressing those involved in differentiation [16] [34]. These factors operate in concert with epigenetic regulators including histone modifiers such as SUV39H1 and DOT1L, as well as DNA methyltransferases, which collectively maintain an open chromatin configuration permissive for pluripotency gene expression [1] [16].
Exogenous signaling pathways provide critical inputs that support pluripotency maintenance and influence differentiation potential. The Wnt/β-catenin signaling pathway promotes self-renewal through stabilization of β-catenin, which interacts with TCF/LEF transcription factors to enhance the expression of pluripotency genes [1]. Conversely, BMP signaling exhibits context-dependent effects, supporting self-renewal in combination with LIF in some contexts while promoting differentiation in others [16]. The TGF-β/Activin A signaling pathway activates SMAD2/3, which regulates Nanog expression and supports pluripotent state maintenance [1] [31]. Additionally, FGF signaling through ERK1/2 supports self-renewal and proliferation, while PI3K/AKT signaling promotes growth and metabolism adapted to pluripotent state requirements [1]. Understanding these pathways is essential not only for maintaining pluripotency but also for directing differentiation into specific lineages for therapeutic applications.
Signaling Dynamics During mRNA Reprogramming
The process of mRNA reprogramming involves dynamic changes in signaling pathways that drive the transition from somatic to pluripotent state. During the initial phase (days 0-4), introduced transcription factors bind to target loci and initiate mesenchymal-to-epithelial transition (MET), accompanied by metabolic shifts from oxidative phosphorylation to glycolysis [34]. The intermediate phase (days 5-12) involves activation of endogenous pluripotency genes and establishment of epigenetic remodeling, with gradual downregulation of somatic cell programs [34]. In the stabilization phase (days 13-18), the cells consolidate the pluripotent state through establishment of autoregulatory loops and chromatin reorganization, ultimately resulting in fully reprogrammed iPSC colonies [1] [34].
The signaling pathways can be visualized as an interconnected network that governs the reprogramming process:
Figure 2: Signaling Pathways in Pluripotency Establishment and Maintenance. This diagram illustrates the core transcription factors (yellow) and key signaling pathways (green) that interact to establish and maintain pluripotency during mRNA reprogramming, with critical cellular processes (red) influenced by these networks.
The Scientist's Toolkit: Research Reagent Solutions
The generation of clinical-grade iPSCs requires carefully selected reagents that comply with GMP standards and support the development of safe, therapeutically applicable cell lines. The following essential materials represent critical components of the clinical-grade iPSC generation workflow:
Table 2: Essential Research Reagents for Clinical-Grade iPSC Generation
| Reagent Category | Specific Examples | Function | Clinical-Grade Considerations |
|---|---|---|---|
| Reprogramming Factors | StemRNA (REPROCELL), Pluristyx mRNA kits | Deliver OSKM factors for cellular reprogramming | Non-integrating, footprint-free, manufactured under GMP conditions [28] [33] |
| Base Media | Pluriton Reprogramming Medium, DMEM/F12-CTS, KnockOut Serum Replacement-CTS | Provide nutritional support for cell growth and reprogramming | Xeno-free, chemically defined, compliant with regulatory standards [31] |
| Growth Factors | human bFGF, human LIF, BDNF, GDNF | Support pluripotency maintenance and direct differentiation | Recombinant human proteins, endotoxin-free, GMP-grade [1] [31] |
| Supplements | N2 Supplement, B27 Supplement, NEAA, GlutaMAX | Enhance cell growth and functionality | Xeno-free formulations, quality-controlled for consistency [31] |
| Extracellular Matrices | Laminin-521, Vitronectin, Recombinant Laminin | Provide substrate for cell attachment and growth | Defined, xeno-free, manufactured under GMP conditions [33] [31] |
| Small Molecule Enhancers | 8-Br-cAMP, Valproic acid, Sodium butyrate, RepSox | Improve reprogramming efficiency and kinetics | Chemical-defined, replace transcription factors in some protocols [1] |
The selection of appropriate reagents is critical for maintaining GMP compliance throughout the iPSC generation process. Particularly important is the use of xeno-free components throughout the entire workflow, as animal-derived products pose risks of immune rejection and transmission of zoonotic pathogens [31]. Additionally, chemically defined formulations ensure lot-to-lot consistency and reduce variability in the reprogramming process and subsequent differentiation protocols [33] [31]. The regulatory documentation provided with clinical-grade reagents, including certificates of analysis and traceability information, is essential for regulatory submissions and quality assurance [33] [32]. Furthermore, compatibility between system components must be verified to ensure optimal performance and reproducibility across multiple cell lines and manufacturing batches [31] [32].
The generation of clinical-grade iPSCs through non-integrative mRNA technology represents a transformative approach in regenerative medicine, combining advanced cellular reprogramming with rigorous quality standards. The adherence to GMP principles and implementation of comprehensive quality control measures throughout the manufacturing process—from donor selection to final cell banking—ensures the safety, identity, potency, and purity of the resulting cell products [30] [33] [31]. The development of footprint-free mRNA reprogramming methods has been particularly instrumental in advancing the field toward clinical applications, eliminating the risk of genomic integration while maintaining high reprogramming efficiency [28] [33] [16].
As the field continues to evolve, ongoing refinements in mRNA technology, differentiation protocols, and quality assurance systems will further enhance the clinical applicability of iPSC-derived therapies. The establishment of standardized frameworks for manufacturing and characterization, coupled with the development of comprehensive iPSC banking initiatives, promises to accelerate the translation of these remarkable cells into routine clinical practice [33] [32] [34]. Through continued adherence to rigorous quality standards and technological innovation, clinical-grade iPSCs are poised to realize their full potential as a transformative therapeutic modality across a broad spectrum of human diseases.
The discovery of induced pluripotent stem cell (iPSC) technology represents a paradigm shift in regenerative medicine and biological research, enabling the reprogramming of somatic cells back to a pluripotent state through the forced expression of specific transcription factors. The original method, employing integrating viral vectors, posed significant clinical risks due to potential genomic alterations and insertional mutagenesis. The advent of non-integrative reprogramming systems, particularly mRNA-based reprogramming, has emerged as a solution, offering a footprint-free method to generate clinical-grade iPSCs. This technique involves the direct delivery of in vitro transcribed mRNA encoding the reprogramming factors, resulting in transient expression without genetic integration [35]. The resulting mRNA-derived iPSCs provide a safe and versatile foundation for disease modeling, drug screening, and the development of autologous cell therapies.
This technical guide details the molecular mechanisms underlying the induction of pluripotency via mRNA technology and provides detailed protocols for the subsequent directed differentiation of these iPSCs into specialized cell types. The entire process is framed within the context of modern non-integrative approaches, which are essential for clinical applications. The core advantages of mRNA reprogramming include its high efficiency, defined genetic footprint, and suitability for industrialized production of stem cells for the clinic [35]. By leveraging this technology, researchers can generate patient-specific iPSC lines that serve as a renewable source for deriving functional somatic cells, thereby powering advanced in vitro models and personalized regenerative treatments.
Molecular Mechanisms of iPSC Induction and Reprogramming
The Molecular Roadmap of Reprogramming
Factor-induced reprogramming is a highly inefficient process, which initially complicated mechanistic studies. However, the isolation of defined intermediate cell populations has enabled a detailed, genome-wide analysis of the journey from a somatic to a pluripotent state. Research has shown that induced pluripotency proceeds through a biphasic process, characterized by two distinct transcriptional waves [36] [11]. The first wave is primarily driven by the expression of c-Myc and Klf4 and occurs within the first few days of factor induction. This phase is marked by the downregulation of somatic genes, the initiation of a mesenchymal-to-epithelial transition (MET), and the activation of processes related to cell proliferation and metabolism [36]. The second wave, which is crucial for the establishment of stable pluripotency, is driven by Oct4, Sox2, and Klf4 and occurs later in the process (after approximately 9 days in mouse models) [36]. This phase activates genes associated with embryonic development and stem cell maintenance.
Cells that ultimately become refractory to reprogramming often successfully activate the first wave but fail to initiate this critical second transcriptional wave [36]. The establishment of a pluripotent epigenetic landscape follows a gradual timeline: bivalent chromatin domains are established progressively after the first wave, while comprehensive changes in DNA methylation, which lock in the pluripotent state, occur predominantly after the second wave, when cells have acquired stable pluripotency [36]. This refined understanding of the molecular sequence allows for the identification of reprogramming roadblocks and the development of strategies to overcome them.
Non-Integrative mRNA Reprogramming Technology
mRNA-based reprogramming is considered the most unambiguously "footprint-free" method for generating iPSCs [35]. This technique involves the repeated transient transfection of somatic cells with synthetic, modified mRNA molecules encoding the key reprogramming factors, typically OCT4, SOX2, KLF4, c-MYC (OSKM). To overcome the innate antiviral immune response triggered by exogenous mRNA, the nucleotides are often modified (e.g., with pseudouridine) and the transcripts are co-delivered with immune-suppressive reagents [16].
The primary molecular advantage of this system is the absence of genomic integration, which eliminates the risk of insertional mutagenesis and produces iPSCs that are more suitable for clinical applications. The transient nature of the mRNA allows for precise control over the timing and dosage of factor expression, leading to more homogeneous reprogramming kinetics. Recent technical improvements have simplified its application, making it a robust and productive method that is increasingly being industrialized for the mass production of human stem cells for the clinic [35]. Compared to other non-integrating methods like Sendai virus or episomal vectors, mRNA reprogramming offers the fastest clearance of the reprogramming factors, resulting in truly footprint-free iPSC clones.
Experimental Protocols for mRNA Reprogramming and Differentiation
Key Research Reagent Solutions
The following table catalogues the essential reagents and their functions required for successful mRNA reprogramming and the subsequent maintenance of iPSCs.
Table 1: Essential Research Reagents for mRNA Reprogramming and iPSC Culture
| Reagent / Material | Function / Explanation |
|---|---|
| Synthetic mRNA (OSKM) | Modified mRNA (e.g., pseudouridine) encoding OCT4, SOX2, KLF4, and c-MYC; the core reprogramming factors that induce pluripotency without genomic integration [35]. |
| Transfection Reagent | A delivery vehicle (e.g., lipid-based) to efficiently introduce mRNA into the cytoplasm of target somatic cells. |
| Immune Suppressant | A small molecule (e.g., B18R) used to temporarily inhibit the innate immune response against transfected mRNA, enhancing cell survival and reprogramming efficiency [16]. |
| Feeder Cells or Defined Matrix | A growth surface; either inactivated mouse embryonic fibroblasts (MEFs) or a defined substrate like Matrigel or laminin-511, to support iPSC attachment and growth. |
| iPSC Culture Medium | A defined medium (e.g., mTeSR1 or E8) containing essential nutrients and growth factors (e.g., FGF2) to maintain pluripotency and self-renewal. |
| Rho-associated Kinase (ROCK) Inhibitor | A small molecule (e.g., Y-27632) used to enhance the survival of single-cell dissociated iPSCs, reducing apoptosis following passaging or thawing. |
Detailed Protocol: mRNA-Based Reprogramming of Human Fibroblasts
This protocol outlines the key steps for reprogramming human dermal fibroblasts (HDFs) using synthetic mRNA.
Step 1: Pre-conditioning and Plating of Somatic Cells
- Culture HDFs in standard fibroblast medium until 70-80% confluent.
- One day before transfection, seed HDFs at a density of 20,000-50,000 cells per well of a 24-well plate coated with a suitable matrix (e.g., gelatin or fibronectin). The goal is to achieve 30-50% confluency at the time of the first transfection.
- Note: Some protocols include a pre-treatment step with the immune suppressant (e.g., 50-100 ng/mL B18R) 12-24 hours before the first mRNA transfection.
Step 2: Daily mRNA Transfection
- Complex the modified mRNA cocktail (containing OSKM factors, plus potentially an mRNA for a fluorescent marker for tracking) with the chosen transfection reagent according to the manufacturer's instructions. A typical total mRNA amount is 0.5-1 µg per well of a 24-well plate.
- Replace the culture medium on the HDFs with fresh medium containing the immune suppressant.
- Gently add the mRNA-transfection reagent complexes to the cells.
- Repeat this daily transfection process for 12-18 consecutive days. The medium is replaced with fresh, immune-suppressant-containing medium prior to each transfection.
Step 3: Transition to iPSC Culture Conditions and Colony Picking
- Between days 7-10, emerging iPSC colonies with a compact, dome-shaped morphology will become visible.
- At this stage, change the culture medium to a defined iPSC maintenance medium (e.g., mTeSR1). The immune suppressant can be omitted from this point forward.
- Between days 14-21, manually pick well-defined, iPSC-like colonies and transfer them to new culture vessels pre-coated with a defined matrix (e.g., Matrigel) containing iPSC medium supplemented with a ROCK inhibitor to enhance survival.
- Expand the clonal lines and perform rigorous characterization, including immunocytochemistry for pluripotency markers (OCT4, NANOG, SSEA-4), karyotyping, and validation of the absence of exogenous reprogramming factors.
Quantitative Data on Reprogramming Dynamics
The following table summarizes key quantitative findings from foundational studies on the dynamics of cellular reprogramming.
Table 2: Quantitative Dynamics of iPSC Reprogramming
| Parameter | Measurement / Finding | Source / Context |
|---|---|---|
| Reprogramming Efficiency | Generally < 3% of starting somatic cells [36]; significantly enhanced by mRNA transfection and intermediate cell sorting. | Studies using OKSM factor expression in fibroblasts. |
| Kinetics of Surface Marker Expression | Thy1− (days 1-2) → SSEA1+ (days 3-5) → Oct4-GFP+ (days 8-10) → Stable iPSC colonies (~day 15) [36]. | Murine embryonic fibroblast (MEF) reprogramming model. |
| Transcriptional Waves | Two distinct waves: 1st wave (days 0-3, Myc/Klf4-driven) and 2nd wave (after day 9, Oct4/Sox2/Klf4-driven) [36]. | Genome-wide analysis of purified intermediate cell populations. |
| Number of Differentially Expressed Genes | Gradual increase, culminating in ~1,500 genes between progressing and refractory populations by day 12 [36]. | Comparison of SSEA1+ (progressing) vs. Thy1+ (refractory) cells. |
Directed Differentiation of iPSCs into Specialized Cells
The true power of iPSC technology lies in the ability to differentiate them into a vast array of functional somatic cells. This process recapitulates embryonic development in vitro by manipulating key signaling pathways.
Principles of Directed Differentiation: Differentiation protocols typically involve a stepwise approach, guiding iPSCs through developmental intermediate stages by adding specific growth factors and small molecules at precise time points. The process is orchestrated by modulating evolutionarily conserved signaling pathways, including BMP, Wnt, Nodal/Activin (TGF-β), and FGF [16]. The initial step often involves the formation of germ layers (ectoderm, mesoderm, endoderm), which are then further specified into target cell types.
Protocol Example: Differentiation into Functional Neurons
- Day 0: Dissociate iPSCs into a single-cell suspension and plate them as a high-density monolayer on a matrix suitable for neural induction (e.g., poly-ornithine/laminin) in neural induction medium containing dual SMAD inhibitors (e.g., SB431542 for TGF-β/Activin inhibition and LDN-193189 for BMP inhibition) and a ROCK inhibitor.
- Days 1-7: Continue culture in neural induction medium with daily medium changes. During this period, cells will form a contiguous sheet of neuroepithelium, characterized by columnar cells. This represents the neural ectoderm stage.
- Days 7-14: Mechanically or enzymatically passage the neural epithelial sheet to form neural progenitor cell (NPC) aggregates or rosettes. Culture in NPC medium containing growth factors like FGF2 and EGF to expand the progenitor population.
- Days 14-35: Induce terminal neuronal differentiation by withdrawing FGF2/EGF and switching to neuronal differentiation medium, often containing brain-derived neurotrophic factor (BDNF), glial cell-derived neurotrophic factor (GDNF), and ascorbic acid. Over the following weeks, progenitors will mature into neurons expressing markers such as TUJ1 (β-III-tubulin) and MAP2.
- Day 35+: Functional maturation can be assessed via electrophysiological measurements (e.g., patch clamping) and immunocytochemistry for synaptic markers (e.g., synapsin).
Signaling Pathways in Directed Differentiation
The following DOT script visualizes the core signaling pathways manipulated during the directed differentiation of iPSCs.
Workflow of mRNA Reprogramming and Differentiation
The complete experimental journey from somatic cell to specialized cell type is depicted in the following workflow diagram.
Applications in Disease Modeling and Drug Development
The combination of mRNA-derived iPSCs and advanced differentiation protocols has opened new frontiers in disease modeling and pharmaceutical research. Patient-specific iPSCs can be generated from individuals with genetic disorders and subsequently differentiated into the cell types affected by the disease. For example, iPSCs from Parkinson's disease patients can be differentiated into dopaminergic neurons, providing a human-relevant model to study disease mechanisms and screen for neuroprotective compounds [16] [11]. These disease-in-a-dish models are particularly valuable for elucidating human-specific phenotypes and molecular pathways that may not be accurately recapitulated in animal models.
Furthermore, the integration of CRISPR-Cas9 gene editing with iPSC technology allows for the creation of isogenic control lines—where the disease-causing mutation is corrected in the patient-derived iPSCs—enabling precise attribution of observed phenotypes to the genetic lesion [16]. This powerful combination is also used to introduce specific mutations into healthy iPSC lines. In drug development, iPSC-derived cells, such as cardiomyocytes and hepatocytes, are increasingly used for high-throughput drug screening and toxicity testing (e.g., cardiotoxicity), providing more predictive human-relevant data earlier in the drug development pipeline [11]. The progression toward more complex 3D models, such as organoids, further enhances the physiological relevance of these systems for modeling tissue-level functions and diseases.
The methodology outlined in this guide—from the footprint-free induction of pluripotency using mRNA technology to the systematic directed differentiation into specialized cells—provides a robust and clinically relevant framework for modern stem cell research. The precise control over reprogramming and differentiation, coupled with a deep understanding of the underlying molecular mechanisms, empowers researchers to create sophisticated human in vitro models. As protocols continue to be refined and integrated with cutting-edge tools like gene editing and machine learning, the potential of mRNA-derived iPSCs to accelerate drug discovery and enable a new generation of personalized regenerative therapies will be fully realized.
Induced pluripotent stem cells (iPSCs) have emerged as a transformative technology in biomedical research and regenerative medicine. This whitepaper provides an in-depth technical analysis of their application in disease modeling, high-throughput drug screening, and personalized therapeutic development, with specific focus on non-integrative mRNA reprogramming methodologies. The integration of advanced gene editing technologies, particularly CRISPR-Cas9, with iPSC platforms has accelerated the development of physiologically relevant human disease models and created new paradigms for drug discovery and cell-based therapies. We present comprehensive experimental protocols, key signaling pathways, and essential research reagents to facilitate implementation of iPSC technology in research and therapeutic development.
Induced pluripotent stem cells (iPSCs) are adult somatic cells that have been reprogrammed to an embryonic-like pluripotent state through the forced expression of specific transcription factors, first demonstrated by Yamanaka and colleagues in 2006 [37] [34]. The original reprogramming method defined a combination of four transcription factors—Oct4, Sox2, Klf4, and c-Myc (OSKM)—as sufficient to revert terminally differentiated cells to pluripotency [37] [38]. This groundbreaking discovery enabled the generation of patient-specific pluripotent cells without the ethical concerns associated with human embryonic stem cells (hESCs) [37] [39].
Table 1: Evolution of iPSC Reprogramming Methods
| Reprogramming Method | Key Factors/Delivery | Advantages | Disadvantages | Clinical Applicability |
|---|---|---|---|---|
| Retroviral/Lentiviral (1st Gen) | OSKM integration | High efficiency | Insertional mutagenesis, tumorigenic risk | Low - research only |
| Sendai Virus (2nd Gen) | OSKM non-integrating viral | Non-integrating, efficient | Viral clearance required, immunogenicity | Medium - requires clearance |
| Non-integrative mRNA (Current) | Modified mRNA OSKM | Footprint-free, controlled expression, high safety | Requires optimized delivery, transient expression | High - GMP compliant |
| Episomal Plasmids | OSKM episomal vectors | Non-integrating, DNA-based | Lower efficiency, potential genomic integration | Medium - requires validation |
| Protein-Based | Recombinant OSKM proteins | Completely non-genetic | Very low efficiency, costly | Medium - technical challenges |
The field has progressively moved toward non-integrative reprogramming methods to address safety concerns associated with viral vector integration, which poses risks of insertional mutagenesis and tumorigenesis [16] [22]. Non-integrative mRNA technology represents a cutting-edge approach that utilizes engineered messenger RNA to transiently express reprogramming factors without genomic integration [16] [28]. This method involves the delivery of modified mRNA sequences that encode the essential transcription factors (typically OCT4, SOX2, KLF4, and c-MYC) to somatic cells [28]. The mRNA constructs are optimized for stability and translational efficiency while minimizing innate immune responses [9] [28]. Unlike viral methods, mRNA reprogramming leaves no genetic footprint in the recipient cells, resulting in genetically stable iPSCs suitable for clinical applications [28] [34].
iPSC-Based Disease Modeling: Methodologies and Applications
Experimental Protocol for Disease Modeling Using mRNA-Derived iPSCs
Step 1: Patient Somatic Cell Collection and Preparation
- Obtain patient-specific somatic cells (typically dermal fibroblasts or peripheral blood mononuclear cells) through minimally invasive procedures [40]
- Culture and expand cells in appropriate media: DMEM with 10% FBS for fibroblasts; RPMI with growth factors for hematopoietic cells
- Validate cell identity through flow cytometry (CD90+ for fibroblasts, CD45+ for blood cells) and ensure absence of microbial contamination
Step 2: mRNA-Based Reprogramming
- Transfect cells with modified mRNA cocktails encoding OSKM factors using lipid nanoparticle delivery systems [28]
- Utilize commercially available mRNA reprogramming kits (e.g., Pluristyx Footprint-Free mRNA technology) or custom-formulated mRNAs [28]
- Perform daily transfections for 12-18 days with careful monitoring of cell morphology changes
- Culture in essential pluripotency-supporting media containing bFGF and TGF-β [39]
Step 3: iPSC Colony Selection and Characterization
- Manually pick emerging iPSC colonies based on embryonic stem cell-like morphology (high nucleus-to-cytoplasm ratio, distinct borders)
- Expand clonal lines and validate pluripotency through:
- Immunocytochemistry for pluripotency markers (OCT4, NANOG, SSEA-4, TRA-1-60)
- RT-PCR analysis of endogenous pluripotency gene expression
- In vitro trilineage differentiation potential (ectoderm, mesoderm, endoderm) [28]
- Karyotype analysis to confirm genomic integrity
Step 4: Directed Differentiation to Disease-Relevant Cell Types
- Differentiate validated iPSCs into target cell types using established protocols:
- Validate differentiated cells through cell-specific markers and functional assays
Step 5: Disease Phenotype Analysis
- Compare patient-specific iPSC-derived cells to isogenic controls or healthy donor lines
- Employ functional assays relevant to the disease pathology (calcium imaging for cardiac arrhythmias, electrophysiology for neuronal disorders, contractility for muscular dystrophies) [39]
- Analyze disease-specific pathological features (protein aggregation, synaptic abnormalities, ion channel dysfunction)
Disease Modeling Applications Across Tissue Types
iPSC technology has been successfully applied to model a wide spectrum of human diseases, providing unprecedented insights into disease mechanisms and progression.
Table 2: iPSC Disease Modeling Applications by Tissue Type
| Disease Category | Specific Diseases Modeled | Key Pathological Features Recapitulated | References |
|---|---|---|---|
| Neurodegenerative | Alzheimer's disease, Parkinson's disease, ALS, Huntington's disease, Spinal muscular atrophy | Aβ and tau pathology, dopaminergic neuron loss, motor neuron degeneration, mHTT aggregation, SMN1 deficiency | [37] [22] [40] |
| Cardiovascular | Long QT syndromes (1,2,7), LEOPARD syndrome, Dilated cardiomyopathy | Action potential prolongation, calcium handling defects, arrhythmogenesis, structural abnormalities | [37] [39] |
| Muscular Dystrophies | Duchenne Muscular Dystrophy (DMD), Becker MD, Facioscapulohumeral MD, Myotonic dystrophy | Dystrophin deficiency, calcium influx abnormalities, myotube alignment defects, nuclear abnormalities | [37] [39] |
| Hepatic | α1-antitrypsin deficiency, Gaucher disease type III | Protein aggregation, enzymatic deficiency, lipid accumulation | [37] |
| Hematological | Fanconi anemia, Thalassemia, Fragile X syndrome | Genomic instability, hemoglobin defects, nucleotide repeat expansion | [37] |
The development of 3D organoid systems has further enhanced the physiological relevance of iPSC-based disease models. These complex, self-organizing structures better recapitulate tissue architecture and cell-cell interactions compared to traditional 2D cultures [16]. For neurological diseases like Alzheimer's, 3D models have enabled the study of Aβ and tau pathology in a more native context, including the observation of tau spreading between connected neurons [40]. Similarly, 3D skeletal muscle models have revealed the importance of the Dystrophin-Associated Protein Complex (DAPC) in myotube alignment and organization, which is disrupted in muscular dystrophies [39].
High-Throughput Drug Screening Using iPSC Platforms
Workflow for iPSC-Based High-Throughput Screening
Diagram 1: iPSC-Based High-Throughput Screening Workflow
Technical Protocol for High-Throughput Screening
Step 1: iPSC Differentiation and Plate Preparation
- Differentiate patient-specific iPSCs into target cell type at scale using optimized protocols
- Seed cells in 384-well or 1536-well plates optimized for high-content imaging
- Ensure uniform cell density and differentiation efficiency across plates through quality control measures
- Allow cells to mature to appropriate developmental stage (e.g., 30-60 days for neurons, 10-15 days for cardiomyocytes)
Step 2: Compound Library Design and Dispensing
- Curate compound libraries (small molecules, FDA-approved drugs, natural products) based on disease mechanism
- Include appropriate controls (positive, negative, vehicle) in each plate
- Use automated liquid handling systems (e.g., Beckman Coulter Biomek) for compound transfer
- Implement concentration-response curves (typically 8-point, 1:3 serial dilutions) for primary screening
Step 3: Assay Implementation and Incubation
- Employ disease-relevant phenotypic assays:
- Calcium-sensitive dyes (Fluo-4) for cardiac arrhythmia models [39]
- Mitochondrial membrane potential dyes (TMRE) for neurodegenerative diseases
- Apoptosis markers (Caspase-3/7 activation) for toxicity screening
- Incubate compounds for appropriate duration (24h-7 days depending on disease model)
Step 4: High-Content Analysis and Data Processing
- Acquire images using automated high-content imaging systems (e.g., PerkinElmer Operetta, ImageXpress Micro)
- Extract multiple parameters simultaneously (morphology, intensity, texture, object count)
- Normalize data to plate controls and apply quality control metrics (Z'-factor > 0.5)
- Use machine learning algorithms for pattern recognition and hit identification
Step 5: Hit Validation and Mechanistic Studies
- Confirm primary hits in dose-response format using fresh compound preparations
- Expand testing to include multiple patient-derived iPSC lines to account for genetic variability
- Investigate mechanism of action through transcriptomics, proteomics, and functional assays
- Evaluate selectivity through counter-screens in relevant cell types
Applications in Drug Discovery and Development
iPSC-based high-throughput screening has been successfully implemented across multiple disease areas, leading to the identification of novel therapeutic candidates and the repositioning of existing drugs:
Neurodegenerative Diseases: Screening of iPSC-derived neurons from Alzheimer's patients identified anti-inflammatory agents (cromolyn) and antiparasitic compounds (avermectins) as potential Aβ-reducing therapies [22]. Similarly, statins were found to modify phosphorylated tau levels through iPSC-based screens [22].
Cardiac Arrhythmias: Patient-specific iPSC-derived cardiomyocytes from Long QT syndrome patients have been used to screen for compounds that correct action potential prolongation, leading to the identification of new anti-arrhythmic candidates [37].
Muscular Dystrophies: DMD iPSC-derived myotubes have enabled screening for compounds that improve cell survival and reduce calcium influx abnormalities, with several candidates advancing to preclinical development [39].
The pharmaceutical industry is increasingly adopting iPSC platforms for toxicity assessment, as they provide more human-relevant data compared to traditional animal models. iPSC-derived cardiomyocytes are now routinely used for assessing cardiotoxicity, while iPSC-derived hepatocytes serve as models for drug-induced liver injury [22] [41].
Personalized Therapeutic Development
iPSC-Based Personalized Therapy Development Pipeline
The application of iPSCs in personalized therapeutic development encompasses two main approaches: patient-specific drug testing and autologous cell therapy.
Table 3: iPSC Applications in Personalized Medicine
| Application | Methodology | Key Advantages | Clinical Stage | Examples |
|---|---|---|---|---|
| Patient-Specific Drug Testing | Derive target cells from patient iPSCs, test drug efficacy and toxicity in vitro | Identifies optimal treatments, avoids adverse reactions, personalized dosing | Clinical implementation | Epilepsy drug selection, chemotherapy sensitivity testing |
| Autologous Cell Therapy | Gene correction of patient iPSCs followed by differentiation and transplantation | Avoids immune rejection, addresses genetic causes, permanent solution | Early clinical trials | Parkinson's disease, macular degeneration, Duchenne Muscular Dystrophy |
| Allogeneic Cell Therapy | HLA-matched iPSC banks, pre-differentiated cell products | Off-the-shelf availability, cost-effective, standardized quality | Advanced clinical trials | Cartilage repair (osteoarthritis), retinal pigment epithelium transplantation |
| Disease Modeling for Target ID | Study disease mechanisms in patient-specific cells, identify novel therapeutic targets | Reveals human-specific pathways, patient-relevant target validation | Preclinical research | Familial Alzheimer's models, rare genetic disorders |
Experimental Protocol for Personalized Therapeutic Development
Step 1: Patient iPSC Generation and Characterization
- Generate clinical-grade iPSCs using GMP-compliant mRNA reprogramming methods [28]
- Comprehensive characterization including identity, purity, potency, and safety profiling
- Whole-genome sequencing to identify disease-causing mutations and potential risk variants
Step 2: Genome Editing for Gene Correction (for genetic disorders)
- Design CRISPR-Cas9 reagents for precise gene correction using non-integrative approaches
- For monogenic disorders (e.g., Duchenne Muscular Dystrophy):
- Design sgRNAs targeting mutation sites
- Use homology-directed repair (HDR) with donor templates for precise correction
- Employ base editing or prime editing for precise nucleotide changes without double-strand breaks [16]
- Validate corrected clones through sequencing and functional assays
Step 3: In Vitro Therapeutic Efficacy Testing
- Differentiate corrected iPSCs into target cell type
- Validate functional correction of disease phenotype:
Step 4: Preclinical Safety and Efficacy Assessment
- Evaluate tumorigenic potential through teratoma formation assays in immunodeficient mice
- Assess functional improvement in disease-relevant animal models
- Perform pharmacokinetic and pharmacodynamic studies for small molecule therapies identified through screening
Step 5: Clinical Translation
- Scale up production under GMP conditions for cell therapies
- Develop optimized delivery methods (scaffolds for tissue engineering, direct injection for cellular therapies)
- Design clinical trials with appropriate endpoints and patient stratification
Signaling Pathways in iPSC Differentiation and Disease Modeling
Diagram 2: Key Signaling Pathways in iPSC Differentiation and Disease Modeling
The Scientist's Toolkit: Essential Research Reagents and Solutions
Successful implementation of iPSC technology requires access to specialized reagents and tools. The following table summarizes essential components for establishing iPSC-based disease modeling, drug screening, and therapeutic development capabilities.
Table 4: Essential Research Reagents for iPSC Applications
| Reagent Category | Specific Products | Key Function | Technical Notes |
|---|---|---|---|
| Reprogramming Kits | Pluristyx Footprint-Free mRNA, CytoTune-iPS Sendai, StemRNA NP | Somatic cell reprogramming to pluripotency | mRNA methods preferred for clinical applications; optimize delivery conditions |
| Cell Culture Media | mTeSR Plus, StemFlex, Essential 8, ReproTeSR | Maintenance of pluripotent state | Feeder-free systems recommended; monitor pluripotency regularly |
| Differentiation Kits | STEMdiff Cardiomyocyte, STEMdiff Neural, PSC-Derived Hepatocyte | Directed differentiation to specific lineages | Optimize for specific applications; validate with functional assays |
| Gene Editing Tools | CRISPR-Cas9 ribonucleoproteins, Base editors, Prime editors | Genetic modification for disease modeling and correction | Use non-integrating methods; validate edits thoroughly |
| Characterization Antibodies | OCT4, NANOG, SSEA-4, TRA-1-60 (pluripotency); βIII-tubulin, cTnT, AFP (differentiation) | Validation of cell identity and differentiation | Use validated panels; include isotype controls |
| Extracellular Matrices | Geltrex, Matrigel, Vitronectin, Laminin-521 | Cell attachment and signaling | Test different matrices for specific applications |
| Small Molecule Modulators | CHIR99021 (Wnt activator), IWP2 (Wnt inhibitor), SB431542 (TGF-β inhibitor), LDN193189 (BMP inhibitor) | Control of differentiation pathways | Optimize concentration and timing for specific protocols |
| Analysis Tools | High-content imaging systems, Multi-electrode arrays, Calcium imaging dyes, Seahorse analyzers | Functional assessment of differentiated cells | Standardize protocols across experiments |
iPSC technology has revolutionized disease modeling, drug screening, and therapeutic development by providing unlimited sources of patient-specific cells that recapitulate disease pathology. The integration of non-integrative mRNA reprogramming methods has addressed critical safety concerns, accelerating clinical translation of iPSC-based applications. Combined with advanced gene editing technologies and high-throughput screening platforms, iPSCs now enable unprecedented opportunities for personalized medicine, from patient-specific drug testing to autologous cell therapies. As the field continues to advance, addressing challenges related to maturation, standardization, and scalability will be essential for fully realizing the potential of this transformative technology in research and clinical applications.
Overcoming Technical Hurdles: Strategies for Enhancing Efficiency and Safety in mRNA Reprogramming
The application of messenger RNA (mRNA) technology for inducing pluripotency represents a paradigm shift in regenerative medicine, offering a non-integrative and controllable strategy for somatic cell reprogramming. Unlike viral vectors that pose risks of insertional mutagenesis, mRNA-based approaches provide a transient, safe method for expressing reprogramming factors while preserving genomic integrity [42]. However, the clinical translation of this technology is hampered by suboptimal reprogramming efficiency, often resulting from inadequate protein expression, imperfect mRNA construct design, suboptimal delivery timing, and unbalanced factor ratios. This technical guide addresses these critical bottlenecks by presenting data-driven optimization strategies framed within the context of advanced mRNA technology for pluripotency research. We focus on three pillars of optimization: mRNA construct engineering, temporal delivery patterns, and factor cocktail composition, providing researchers with a comprehensive framework for enhancing reprogramming outcomes.
mRNA Construct Design: Engineering for Enhanced Expression and Reduced Immunogenicity
The foundational element of successful reprogramming lies in the strategic design of mRNA constructs. Optimal design significantly enhances translation efficiency, prolongs protein expression, and minimizes innate immune responses that can derail reprogramming.
Nucleoside Modifications and Capping Strategies
Incorporation of modified ribonucleosides is critical for evading cellular antiviral defenses. Research demonstrates that complete substitution of cytidine with 5-methylcytidine (5mC) and uridine with pseudouridine (ψ) markedly improves cell viability and increases ectopic protein expression [43]. These modifications dramatically attenuate interferon signaling by reducing activation of pattern recognition receptors like RIG-I and PKR [43].
The capping strategy equally influences translational efficiency. Use of a virus-derived capping enzyme instead of cap analogs ensures 100% proper cap orientation, creating a cap1 structure found in higher eukaryotes with superior translation efficiency compared to other methods [7]. This approach, combined with optimized 5' and 3' untranslated regions (UTRs), can boost protein expression levels and duration.
Codon Optimization and Sequence Engineering
Recent advances in deep learning have revolutionized codon optimization. The RiboDecode framework demonstrates how generative AI can explore vast sequence spaces to design mRNA codon sequences with enhanced translational characteristics [19]. Unlike traditional rule-based approaches like codon adaptation index (CAI), RiboDecode directly learns from large-scale ribosome profiling data to predict translation levels, resulting in mRNA constructs with substantially improved protein expression [19].
Table 1: Key mRNA Construct Modifications and Their Functional Impact
| Modification Type | Specific Approach | Functional Impact | Experimental Evidence |
|---|---|---|---|
| Nucleoside Modification | ψ and 5mC incorporation | Reduces innate immune activation; increases translational efficiency | 50-90% transfection efficiency across human cell types [43] |
| Capping Strategy | Virus-derived capping enzyme | 100% proper cap orientation; enhanced translation | Superior to cap analogs; creates eukaryotic cap1 structure [7] |
| Codon Optimization | RiboDecode deep learning framework | Enhanced translation efficiency; exploration of novel sequence space | Substantial improvements in protein expression vs. traditional methods [19] |
| UTR Engineering | Alpha-globin 3' UTR with strong Kozak sequence | Improved translational initiation and mRNA stability | Sustained protein expression for several days [43] |
Delivery Timing and Regimen: The Temporal Dimension of Reprogramming
The transient nature of mRNA-mediated protein expression necessitates precise delivery timing to maintain sustained factor levels throughout the reprogramming process.
Frequency and Duration Optimization
Daily transfection of modified mRNA has been established as an effective regimen for maintaining sufficient reprogramming factor levels. Research shows that repeated administration of synthetic mRNAs incorporating nucleoside modifications enables reprogramming of differentiated human cells with efficiencies substantially superior to established viral protocols [43]. The optimal duration typically spans 10-18 days, with pluripotency markers emerging as early as day 8 under optimized conditions.
The inclusion of B18R protein, a vaccinia virus decoy receptor for Type I interferons, in the media further supports repeated transfections by mitigating residual interferon responses [43]. This combination allows for sustained high-level protein expression without substantial cytotoxicity, enabling the extended expression window required for complete epigenetic remodeling.
Staggered and Sequential Delivery Approaches
Emerging evidence suggests that delivering reprogramming factors in a staggered sequence rather than as a single cocktail may enhance efficiency. This approach mimics the natural embryonic development process where specific factors are expressed in temporal waves. While optimal sequences require further validation, preliminary data indicate that initiating with OS (OCT4 and SOX2) followed by KM (KLF4 and c-MYC) after 3-5 days may yield superior results compared to concurrent delivery.
Factor Cocktail Ratios: Balancing Reprogramming and Pluripotency
The composition and stoichiometry of reprogramming factors significantly influence both the efficiency and quality of resulting induced pluripotent stem cells (iPSCs).
Core Factor Optimization
The canonical OSKM (OCT4, SOX2, KLF4, c-MYC) cocktail remains the foundation for most reprogramming protocols, but the optimal ratio varies by cell type and delivery method. Research indicates that fine-tuning the relative proportions of these factors can dramatically impact efficiency. For instance, certain cell types with endogenous expression of specific factors may require adjusted ratios [44].
Alternative cocktails including OSNL (OCT4, SOX2, NANOG, LIN28) have shown promise, with some studies demonstrating that a six-factor combination (OSKMNL) can increase reprogramming efficiency by 10-fold compared to simpler combinations [44]. This enhanced cocktail has proven particularly valuable for challenging cell sources, including senescent cells from aged donors.
Supplemental Enhancers and Small Molecules
The addition of epigenetic modifiers and other small molecules can substantially boost reprogramming efficiency. Compounds such as valproic acid (histone deacetylase inhibitor), sodium butyrate, and 5-azacytidine (DNA methyltransferase inhibitor) have demonstrated the ability to enhance reprogramming efficiency by 15-51 times in some systems [44]. These compounds work by remodeling the epigenetic landscape to facilitate the transition to pluripotency.
Table 2: Reprogramming Enhancement Compounds and Their Mechanisms
| Compound Category | Specific Examples | Mechanism of Action | Impact on Efficiency |
|---|---|---|---|
| Histone Deacetylase Inhibitors | Valproic acid, Sodium butyrate | Opens chromatin structure; facilitates epigenetic remodeling | Can replace oncogene c-MYC or KLF4; 15-51x improvement [44] |
| DNA Methyltransferase Inhibitors | 5-Azacytidine | Reduces DNA methylation barriers | Facilitates transition to pluripotent state [44] |
| Histone Demethylase Inhibitors | Parnate | Modifies histone methylation patterns | Enables reprogramming with just OCT4 and KLF4 [44] |
| Metabolic Cofactors | Vitamin C | Alleviates cell senescence; promotes demethylation | Increases colony numbers when combined with valproic acid [44] |
Integrated Workflow: From Design to Validation
A systematic, integrated approach combining optimized constructs, timing, and ratios delivers the most robust reprogramming outcomes. The following workflow visualization encapsulates the key decision points and their relationships in establishing an optimized mRNA reprogramming protocol:
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of optimized mRNA reprogramming requires access to specific, high-quality reagents. The following table details essential materials and their functions:
Table 3: Essential Research Reagents for mRNA Reprogramming
| Reagent Category | Specific Examples | Function/Purpose | Technical Notes |
|---|---|---|---|
| Nucleoside Modifications | Pseudouridine-5'-TP, 5-Methylcytidine-5'-TP | Reduces innate immune recognition | Complete substitution of uridine and cytidine [43] |
| Capping System | Vaccinia virus capping enzyme, 2'-O-Methyltransferase | Creates cap1 structure; enhances translation | Superior to cap analogs; provides 100% proper orientation [7] |
| Interferon Inhibitor | B18R recombinant protein | Blocks interferon signaling; enables repeated transfections | Critical for sustained expression over multi-day protocol [43] |
| Transfection Reagent | Polyethylenimine (PEI) | Facilitates mRNA delivery; cationic complexing agent | Superior to other methods for repeated transfections [7] [43] |
| Reprogramming Factors | OCT4, SOX2, KLF4, c-MYC mRNA | Core reprogramming transcription factors | Modified mRNAs with enhanced stability and translation [44] [43] |
| Small Molecule Enhancers | Valproic acid, Sodium butyrate, Vitamin C | Epigenetic modifiers; senescence alleviation | Significantly boosts efficiency; enables difficult reprogramming [44] |
Optimizing mRNA-based reprogramming requires a multifaceted approach addressing construct design, delivery parameters, and factor composition in an integrated manner. The strategies outlined in this technical guide provide a roadmap for significantly enhancing reprogramming efficiency while maintaining the critical safety advantages of non-integrative approaches. As mRNA technology continues to evolve, with advancements in AI-driven sequence design and novel delivery systems, researchers are positioned to overcome current limitations in efficiency. This progress will accelerate the clinical translation of iPSC technologies, enabling new regenerative medicine applications while respecting the stringent safety requirements of human therapeutics. The future of pluripotency research lies in the continued refinement of these mRNA-based approaches, moving toward a standardized, efficient, and clinically viable reprogramming methodology.
The development of non-integrative mRNA technology represents a pivotal advancement in pluripotency research and regenerative medicine. Unlike traditional gene therapy approaches that risk insertional mutagenesis, synthetic mRNA offers a transient, non-integrative method for delivering reprogramming factors to generate induced pluripotent stem cells (iPSCs) [9] [4]. A primary challenge in this field is the innate immune response triggered by exogenous RNA, which can lead to severe cytotoxicity, impede reprogramming efficiency, and potentially compromise therapeutic outcomes [4] [45]. This technical guide examines two synergistic strategies for managing these immune responses: the incorporation of nucleotide modifications to make mRNA less recognizable to the innate immune system, and the controlled use of immunosuppressive agents to mitigate residual inflammation. Within the context of pluripotency research, mastering these approaches is essential for enhancing the safety and efficiency of iPSC generation, thereby accelerating their application in disease modeling and cell-based therapies [4] [16].
Nucleotide Modifications to Mitigate Immune Sensing
The innate immune system possesses a sophisticated array of pattern recognition receptors (PRRs) designed to detect foreign RNA, a common signature of viral infection. In vitro transcribed (IVT) mRNA can inadvertently activate these pathways, leading to the production of type I interferons and pro-inflammatory cytokines, which in turn can inhibit translation and trigger apoptosis [45] [46].
Key Modifications and Their Mechanisms
Extensive research has identified specific nucleotide modifications that enable synthetic mRNA to evade immune detection while enhancing its stability and translational capacity.
Table 1: Key Nucleotide Modifications and Their Immunomodulatory Effects
| Modification | Effect on Innate Immune Recognition | Impact on Translation & Stability | Key Supporting Research |
|---|---|---|---|
| Pseudouridine (Ψ) | Reduces activation of TLR7/8 and other intracellular sensors like PKR [45] [46]. | Improves translational efficiency and mRNA stability [45]. | Karikó et al. (2005, 2008); [46] |
| N1-methylpseudouridine (m1Ψ) | Superior immuno-evasion compared to Ψ; used in COVID-19 mRNA vaccines [45]. | Further enhances protein yield [45]. | Moderna/Pfizer-BioNTech COVID-19 vaccines [45] |
| 5-methylcytidine (m5C) | Decreases immune stimulation; often used in combination with Ψ [4] [46]. | Contributes to mRNA stability [46]. | [46] |
The seminal work by Karikó and Weissman demonstrated that replacing uridine with pseudouridine allows mRNA to be recognized as "self" rather than "non-self," drastically reducing interferon signaling [45]. This foundational discovery enabled the clinical success of mRNA vaccines, which employ m1Ψ to achieve high levels of antigen expression with minimal reactogenicity [45]. Beyond uridine derivatives, modifying cytidine to 5-methylcytidine also contributes to dampening the immune response, a finding corroborated in studies on small nuclear RNA (snRNA) and small nucleolar RNA (snoRNA) analogs [46]. The combination of these modifications has a synergistic effect, further diminishing the activation of sensors such as protein kinase R (PKR), which can phosphorylate translation initiation factor eIF2α to shut down global protein synthesis [46].
Experimental Protocols for Validation
To assess the efficacy of nucleotide modifications in suppressing immune activation, researchers employ a suite of in vitro and in vivo assays.
In Vitro Transcription of Modified mRNA:
- Template Preparation: Generate DNA templates via PCR, incorporating a T7 RNA polymerase promoter sequence and a poly(T) tail for a defined poly(A) tail length [4].
- IVT Reaction: Perform transcription using T7 RNA polymerase. To produce modified RNA, replace canonical NTPs with modified equivalents (e.g., ΨTP or m5CTP) in the reaction mix. Include a cap analog (e.g., m32.2.7G[5′]ppp[5′]G) for 5' capping during transcription [4] [46].
- Purification and Processing: Purify the RNA product. Treat with DNase I to remove template DNA and with phosphatase to remove 5'-triphosphates, which are themselves immunostimulatory. Further purification can be achieved via HPLC or cellulose-based methods to remove double-stranded RNA (dsRNA) contaminants, potent inducers of innate immunity [45] [46].
Assessing Immune Activation:
- Whole-Transcriptome Analysis (RNA-Seq): Transfert cells (e.g., human fibroblasts or MCF-7 cells) with modified or unmodified RNA analogs. Extract total RNA and perform RNA sequencing. Analyze differential expression of interferon-stimulated genes (ISGs) such as OAS1, IFIT1, and MX1 to quantify the interferon response [47] [46].
- Protein-Level Analysis: Use Western blotting or ELISA to measure phosphorylation of PKR and its substrate eIF2α, as well as secretion of cytokines like IFN-β and IL-6 [46].
- shRNA Knockdown Validation: To confirm the role of specific receptors, use stable cell lines expressing shRNA targeting PRRs like PKR (EIF2AK2) or RIG-I (DDX58). Repeating the transfection experiments in these knockdown backgrounds can show a blunted immune response to unmodified RNA [46].
Diagram 1: mRNA Modification and Immune Evasion
Immunosuppressive Agents in mRNA-Based Workflows
Despite nucleotide modifications, the lipid nanoparticle delivery vehicle and residual RNA sensing can still provoke a significant innate immune response [45] [48]. This is particularly critical in sensitive applications like cellular reprogramming, where prolonged cell health is essential.
Application in iPSC Generation
The process of generating iPSCs via synthetic mRNA requires daily transfections over 1-2 weeks, creating a sustained risk of interferon-induced cell death [4]. To ensure cell survival and improve reprogramming efficiency, the use of immunosuppressive agents is a standard practice.
Key Agent: B18R Protein
- Function: The B18R protein is a recombinant version of the vaccinia virus-encoded type I interferon receptor. It acts as a decoy receptor, binding to and neutralizing IFN-α, IFN-β, and IFN-ω in the cell culture medium, thereby preventing the activation of the interferon signaling cascade [4].
- Protocol: The reprogramming medium is supplemented with B18R throughout the mRNA transfection period. Its inclusion is considered essential for successful footprint-free iPSC generation using both standard synthetic mRNA and self-replicating RNA (srRNA) methods [4].
Table 2: Immunosuppressive Agents in mRNA Reprogramming
| Agent | Mechanism of Action | Application in mRNA Workflow | Considerations |
|---|---|---|---|
| B18R Protein | Binds and neutralizes type I interferons (IFN-α/β) in the extracellular medium [4]. | Added to culture medium during prolonged mRNA transfection (e.g., iPSC reprogramming) [4]. | Critical for preventing IFN-induced cell death; used transiently. |
| Small Molecule Inhibitors | Target intracellular signaling nodes (e.g., JAK/STAT pathway) [49]. | Potential use for suppressing persistent immune activation. | Not commonly reported in standard protocols; requires careful dosing. |
Integration in Non-Integrative mRNA Reprogramming
The combination of nucleotide-modified mRNA and transient immunosuppression forms the cornerstone of modern, safe iPSC generation.
Experimental Workflow for iPSC Generation
The following protocol outlines the key steps for generating footprint-free iPSCs, integrating the strategies for immune response management discussed in this guide.
Diagram 2: Non-Integrative iPSC Generation Workflow
Detailed Protocol:
- mRNA Production: Synthesize mRNAs encoding the reprogramming factors (typically Oct4, Sox2, Klf4, c-Myc, and optionally Lin28) via in vitro transcription, incorporating pseudouridine and 5-methylcytidine in place of their canonical counterparts. Include a Cap1 structure and a poly(A) tail to enhance translation and stability [4].
- Cell Transfection: Seed human fibroblasts (e.g., neonatal foreskin fibroblasts) and begin daily transfections with the modified mRNA cocktail. This process typically continues for 12-16 days [4].
- Immunosuppression: Supplement the culture medium with B18R protein for the duration of the transfection period to neutralize any type I interferon response [4].
- Colony Picking and Expansion: Monitor for the emergence of embryonic stem cell-like colonies. Pick and expand putative iPSC clones for further characterization [4].
- Validation of Pluripotency: Confirm the pluripotent state through:
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for mRNA-Based Reprogramming and Immune Management
| Reagent / Material | Function | Example |
|---|---|---|
| Modified Nucleotides | Incorporated into IVT mRNA to reduce immunogenicity and enhance translation. | Pseudouridine-5'-triphosphate (ΨTP), 5-methylcytidine-5'-triphosphate (m5CTP) [4] [46]. |
| Cap Analog | Provides 5' cap structure for improved translation and reduced immune recognition by IFIT proteins. | CleanCap or m32.2.7G[5′]ppp[5′]G analog [4] [45]. |
| B18R Protein | Neutralizes type I interferons in cell culture medium to prevent IFN-mediated cell death. | Recombinant vaccinia virus B18R protein [4]. |
| Ionizable Lipid Nanoparticles (iLNPs) | Efficiently delivers mRNA into the cell cytoplasm; also provides adjuvant activity. | LNPs containing SM-102 or ALC-0315 [45]. |
| T7 RNA Polymerase | Enzyme for in vitro transcription of mRNA from a DNA template. | High-yield T7 RNA polymerase kits [4]. |
The strategic integration of nucleotide modifications and targeted immunosuppressive agents is critical for harnessing the full potential of non-integrative mRNA technology in pluripotency research. Modifications like pseudouridine and 5-methylcytidine directly address the problem of innate immune recognition at the molecular level, while reagents like the B18R protein provide a crucial safety net by managing residual inflammatory responses in cell culture. As the field advances, further refinement of these strategies—such as the development of novel modified nucleotides and biodegradable lipid nanoparticles—will continue to improve the safety, efficiency, and clinical applicability of iPSC-based therapies, paving the way for a new era in regenerative medicine.
The advent of non-integrative mRNA technology for inducing pluripotency has revolutionized the generation of induced pluripotent stem cells (iPSCs) by offering a safer alternative to traditional viral methods. This technology utilizes synthetic mRNA to transiently express the reprogramming factors, eliminating the risk of genomic integration and subsequent insertional mutagenesis [16] [50]. However, despite this enhanced safety profile, the risks of genomic instability and tumorigenicity remain critical concerns that necessitate rigorous purity assessment and characterization of the resulting iPSC lines. Ensuring the integrity of iPSCs is paramount for their reliable application in disease modeling, drug discovery, and regenerative medicine [50].
This technical guide provides an in-depth framework for the comprehensive evaluation of iPSC lines, with a specific focus on methodologies and assays to ensure genomic stability and prevent tumorigenic potential. The protocols and strategies outlined herein are designed to be integrated seamlessly with the non-integrative mRNA reprogramming workflow, providing researchers with a standardized approach for quality control.
Critical Risks in iPSC Generation and Culture
Even with non-integrative reprogramming methods, iPSCs are susceptible to acquiring genetic and epigenetic abnormalities during the reprogramming process and subsequent long-term culture. These aberrations can confer a selective growth advantage to certain clones, a phenomenon known as "culture adaptation" [50]. Furthermore, incomplete reprogramming or residual expression of reprogramming factors can lead to heterogeneous cell populations with increased tumorigenic potential. The risk of tumor formation, particularly teratomas, from residual undifferentiated iPSCs is a primary safety hurdle for clinical applications [50]. Therefore, a multi-parameter assessment strategy is essential to mitigate these risks.
Comprehensive Characterization of iPSC Lines
A robust characterization strategy must confirm the successful establishment of pluripotency and, simultaneously, screen for potential safety hazards. The following sections detail the essential components of this strategy.
Pluripotency Assessment
Confirmation of pluripotency is a foundational step. This involves verifying the expression of key pluripotency markers and the functional capacity to differentiate into all three germ layers.
Table 1: Key Pluripotency Markers for iPSC Characterization
| Marker Type | Marker Genes | Detection Method | Significance |
|---|---|---|---|
| Core Transcription Factors | OCT4 (POU5F1), NANOG, SOX2 [51] [52] | qPCR, Immunocytochemistry | Master regulators of the pluripotency network; essential for self-renewal. |
| Surface Antigens | SSEA-4, TRA-1-60, TRA-1-81 [53] [50] | Flow Cytometry, Immunofluorescence | Characteristic cell surface markers of undifferentiated human pluripotent stem cells. |
| Functional Capacity | In Vitro Trilineage Differentiation | Directed Differentiation | Confirms potential to form ectoderm, mesoderm, and endoderm lineages. |
Experimental Protocol: qPCR Analysis for Pluripotency Markers
- RNA Extraction: Isolate total RNA from a confluent well of a 6-well plate using a column-based kit with DNase I treatment to remove genomic DNA contamination.
- cDNA Synthesis: Convert 1-2 µg of RNA into complementary DNA (cDNA) using a reverse transcriptase enzyme and oligo(dT) or random hexamer primers [52].
- Quantitative PCR: Perform qPCR reactions in triplicate using gene-specific primers for core pluripotency markers (e.g., OCT4, NANOG, SOX2). Include reference genes (e.g., GAPDH, ACTB) for normalization.
- Data Analysis: Calculate relative gene expression using the ΔΔCt method. Compare expression levels to a validated positive control (e.g., an established pluripotent stem cell line) and a negative control (e.g., the parental somatic cells) [52].
Genomic Stability Assessment
Karyotypic abnormalities are a major risk in iPSC culture. Regular monitoring for genetic changes is non-negotiable for both research and clinical grades.
Table 2: Genomic Stability Monitoring Methods
| Method | Resolution | Target Anomaly | Throughput |
|---|---|---|---|
| Karyotyping (G-banding) | ~5-10 Mb | Aneuploidies, large translocations/inversions | Low |
| Array Comparative Genomic Hybridization (aCGH) | ~50-100 kb | Copy Number Variations (CNVs) | Medium-High |
| Error-Corrected Next-Generation Sequencing (ecNGS) | Single Nucleotide | Point mutations, small indels | High |
Advanced techniques like error-corrected Next-Generation Sequencing (ecNGS) are emerging as powerful tools for detecting low-frequency mutations that would be missed by other methods. ecNGS involves sequencing both strands of DNA independently, enabling the bioinformatic identification and filtering of sequencing errors to reveal true, rare mutations with high sensitivity (as low as 1 in 10^7 bases) [54]. This is particularly valuable for identifying mutations in genes associated with cancer.
Tumorigenicity Risk Assessment
The tumorigenic potential of an iPSC line can arise from residual undifferentiated cells or from acquired oncogenic mutations.
Experimental Protocol: Teratoma Assay
- Procedure: Harvest undifferentiated iPSCs, resuspend in a cold Basement Membrane Matrix (e.g., Matrigel), and inject intramuscularly or subcutaneously into immunodeficient mice (e.g., NSG mice). A typical injection contains 1-5 million cells.
- Endpoint Analysis: Tumors are harvested 8-16 weeks post-injection or when they reach a predetermined size (e.g., 1.5 cm diameter). The tissue is fixed, sectioned, and stained with hematoxylin and eosin (H&E).
- Assessment: A qualified histopathologist examines the sections for the presence of differentiated tissues derived from all three embryonic germ layers (ectoderm, mesoderm, and endoderm). The absence of undifferentiated cells and malignant components is critical for de-risking the cell line [50].
In Vitro Surrogate Assays
While the teratoma assay is considered a gold standard, it is time-consuming, expensive, and raises ethical concerns. Flow cytometric analysis for pluripotency surface markers (e.g., SSEA-4, TRA-1-60) provides a rapid, quantitative measure of the undifferentiated cell fraction in a population, which can be correlated with tumorigenic risk [50].
An Integrated Workflow for iPSC Characterization
The following diagram illustrates the sequential, multi-parameter workflow for the comprehensive characterization of iPSC lines generated via non-integrative mRNA reprogramming.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for iPSC Characterization
| Reagent / Kit | Function | Application Note |
|---|---|---|
| Directed Trilineage Differentiation Kit [51] | Induces differentiation into definitive endoderm, mesoderm, and ectoderm lineages. | Used for functional validation of pluripotency. Prefer kits compatible with defined, serum-free media. |
| qPCR Pluripotency Marker Kit [52] | Contains pre-validated primers/probes for core pluripotency genes (e.g., NANOG, OCT4, SOX2). | Ensures specificity and reproducibility in gene expression analysis. Includes housekeeping genes for normalization. |
| Flow Cytometry Antibody Panel [51] [50] | Antibodies against surface markers (SSEA-4, TRA-1-60) and intracellular factors (OCT3/4). | Enables quantitative assessment of purity and pluripotency marker expression at the single-cell level. |
| aCGH or SNP Microarray Kit | Genome-wide screening for copy number variations (CNVs). | A higher-resolution alternative to karyotyping for monitoring genomic integrity. |
| ecNGS Library Prep Kit [54] | Prepares DNA libraries for error-corrected sequencing to detect low-frequency mutations. | Critical for identifying oncogenic mutations; requires specialized bioinformatics analysis. |
| Basement Membrane Matrix | Provides a 3D substrate for cell injection in the teratoma assay. | Mimics the in vivo environment for tumor formation and differentiation. |
Advanced Technologies and Future Perspectives
The field of iPSC characterization is being enhanced by the integration of advanced computational and molecular tools. Machine learning algorithms, such as the "hiPSCore" scoring system, are being developed to classify pluripotent and differentiated cells accurately based on a refined set of marker genes, reducing time and subjectivity in quality control [51]. Furthermore, CRISPR-Cas9 gene editing is used not only for therapeutic correction but also to engineer "hypoimmunogenic" iPSC lines by knocking out HLA genes, thereby reducing the risk of immune rejection upon transplantation [16] [50]. As the field progresses, the combination of non-integrative reprogramming with these rigorous and evolving characterization standards will pave the way for safer clinical translations of iPSC-based therapies.
The emergence of non-integrative mRNA technology has revolutionized pluripotency research and regenerative medicine, offering unprecedented control over cellular reprogramming. Unlike traditional gene therapy approaches that permanently alter the host genome, mRNA-based methods provide transient, precise expression of reprogramming factors with minimal risk of genomic integration. This technical guide examines the critical intersection of mRNA precision dosing and protein expression kinetics as it applies to the generation of induced pluripotent stem cells (iPSCs). We explore fundamental principles of mRNA amplification dynamics, delivery system optimization, and kinetic modeling that enable researchers to achieve consistent, high-quality cellular reprogramming outcomes. Through systematic analysis of current literature and experimental evidence, this whitepaper provides a comprehensive framework for implementing controlled protein expression protocols in pluripotency research applications.
The success of somatic cell reprogramming using mRNA technology hinges on precisely controlling the temporal expression and stoichiometric ratios of key pluripotency factors. The foundational work of Takahashi and Yamanaka demonstrated that overexpression of specific transcription factors—primarily OCT4, SOX2, KLF4, and c-MYC (OSKM)—can reprogram mature, differentiated cells into induced pluripotent stem cells (iPSCs) [34]. However, the transition from viral delivery methods to non-integrative mRNA platforms introduces unique challenges in maintaining optimal expression dynamics for efficient reprogramming.
Traditional viral methods provide sustained expression but carry significant safety concerns due to genomic integration [16]. In contrast, mRNA-based delivery offers a transient, non-integrative alternative but requires precise dosing control to maintain therapeutic protein levels within the optimal window for cellular reprogramming. The fundamental characteristic of mRNA technology involves an amplification process wherein a single mRNA molecule can produce 10³-10⁶ protein copies depending on construct optimization and cellular context [55]. This amplification creates both opportunities and constraints that must be carefully navigated in reprogramming applications.
Table 1: Key Advantages of mRNA Technology for Pluripotency Research
| Feature | Technical Benefit | Impact on Reprogramming |
|---|---|---|
| Non-integrative | No genomic modification | Enhanced safety profile; reduced tumorigenic risk |
| Transient Expression | Controlled protein duration | Prevents sustained oncogene expression |
| Precision Dosing | Tunable protein levels | Optimal factor stoichiometry for reprogramming |
| Rapid Onset | Protein expression within 2-6 hours | Accelerates initiation of reprogramming cascade |
| Chemical Modifications | Enhanced stability and translation | Extended functional protein expression |
Fundamental Principles of mRNA Expression Kinetics
Amplification Dynamics and Protein Yield
The therapeutic efficacy of mRNA in reprogramming applications depends on its ability to hijack cellular protein synthesis machinery, with each mRNA molecule potentially undergoing hundreds to thousands of translation cycles before degradation [55]. Translation efficiency represents a critical factor in determining reprogramming outcomes, with optimized mRNA constructs incorporating 5' and 3' untranslated regions, modified nucleotides, and codon optimization achieving translation rates of 10-100 proteins per mRNA per minute [55]. The amplification effect is particularly relevant for pluripotency factor expression, where precise ratios of transcription factors directly influence reprogramming efficiency and iPSC quality.
Chemical modifications and optimized UTR sequences extend mRNA half-life from minutes for unmodified constructs to 24-72 hours for optimized versions, directly correlating with total protein output [55]. This extended half-life is crucial for reprogramming applications, as the process requires sustained expression of pluripotency factors over several days to weeks to complete the epigenetic transition to pluripotency.
Characteristic Expression Kinetics Profile
Following delivery, therapeutic mRNA exhibits a characteristic temporal expression profile that represents a fundamental consideration for reprogramming protocol design. This pattern remains consistent across different target proteins and delivery routes [55]:
- Rapid Onset: Initial protein detection within 2-6 hours post-administration
- Peak Expression: Maximum protein levels achieved at 24-48 hours
- Exponential Decline: Progressive reduction over 7-14 days
This kinetic profile necessitates repeated dosing for applications requiring sustained protein expression, such as cellular reprogramming, where the process typically requires 2-4 weeks to complete. The frequency and magnitude of these dosing intervals directly impact reprogramming efficiency and the quality of resulting iPSC colonies.
Table 2: mRNA Expression Kinetics and Implications for Reprogramming
| Kinetic Phase | Timeframe | Reprogramming Implications | Optimization Strategies |
|---|---|---|---|
| Rapid Onset | 2-6 hours | Quick initiation of reprogramming cascade | Optimize delivery efficiency; cell synchronization |
| Peak Expression | 24-48 hours | Critical period for reprogramming initiation | Balance factor stoichiometry; avoid cytotoxicity |
| Decline Phase | 7-14 days | Determines dosing frequency | Modify nucleosides; optimize UTRs |
| Total Duration | 7-14 days | Multiple doses needed for complete reprogramming | Schedule repeated administrations (every 24-72 hours) |
Delivery Systems and Dosing Control for Reprogramming
Lipid Nanoparticle Platforms
Current clinical mRNA therapeutics predominantly utilize ionizable lipid nanoparticles (LNPs) as delivery vehicles. The standard composition includes [55]:
- Ionizable lipids (35-50%) for pH-dependent membrane fusion and endosomal escape
- Phospholipids (10-15%) for membrane stability and biocompatibility
- Cholesterol (25-40%) for membrane fluidity modulation
- PEG-lipids (1-3%) for steric stabilization and biodistribution influence
While LNPs have demonstrated clinical success, they exhibit inherent limitations for precision dosing applications in reprogramming, including preferential hepatic accumulation, limited tissue targeting, and batch-to-batch variability in delivery efficiency [55]. Recent innovations include SORT nanoparticles that tune mRNA release based on modulation of internal charge, thereby facilitating delivery to specific tissue types [55].
Advanced Formulation Strategies
The development of organ-specific LNP formulations through lipid modification, targeting ligands, and surface functionalization holds promise for reducing off-target effects and enhancing dosing precision in reprogramming applications [55]. Incorporation of biodegradable polymers, hydrogels, and implantable devices may enable sustained mRNA release and more predictable protein expression kinetics [55]. These advanced delivery strategies are particularly relevant for in vivo reprogramming approaches, where precise spatial control of pluripotency factor expression is essential for preventing teratoma formation and directing specific cellular transitions.
Experimental Protocols for mRNA Reprogramming
mRNA Reprogramming Factor Design
The core reprogramming factors require careful sequence optimization for mRNA-based expression. The most common factors include [1] [34]:
- OCT4 (POU class 5 homeobox 1): Master regulator of pluripotency
- SOX2 (SRY-box transcription factor 2): Essential for self-renewal
- KLF4 (Kruppel-like factor 4): Facilitates epigenetic remodeling
- c-MYC (Myelocytomatosis oncogene): Enhances proliferation and efficiency
Alternative factor combinations have been explored to optimize safety and efficiency. The OSNL combination (OCT4, SOX2, NANOG, LIN28) represents a viable alternative to the traditional OSKM factors, with NANOG functioning as an essential factor for maintaining pluripotency and LIN28 accelerating cell proliferation similar to c-MYC [34]. Studies have demonstrated that the specific ratio of SOX2 to OCT4 during reprogramming is critical, with improper ratios significantly reducing reprogramming efficiency and iPSC quality [34].
Footprint-Free mRNA Reprogramming Methodology
Pluristyx's proprietary approach exemplifies current best practices in mRNA reprogramming, utilizing non-modified mRNA constructs that efficiently reprogram adult cells into genetically stable iPSCs [28]. This platform utilizes mRNA sequences with non-modified nucleotides organized into stable structures to provide long-lasting expression of transcription factors, enabling footprint-free reprogramming without genomic integration [28].
Detailed Protocol:
- Cell Preparation: Plate appropriate somatic cells (typically fibroblasts or peripheral blood mononuclear cells) at optimal density
- mRNA Transfection: Complex modified mRNA reprogramming factors with appropriate transfection reagent
- Dosing Schedule: Transfect cells daily for 14-21 days with precise factor ratios
- Media Optimization: Use specialized media supporting both transfection and pluripotent cell growth
- Colony Selection: Identify and pick emerging iPSC colonies based on morphological criteria
- Quality Validation: Rigorously test resulting iPSCs for pluripotency markers and differentiation potential
This protocol emphasizes the critical importance of dosing precision throughout the reprogramming process, as inconsistent factor expression can lead to partial reprogramming or aberrant iPSC phenotypes.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Research Reagent Solutions for mRNA Reprogramming
| Reagent Category | Specific Examples | Function in Reprogramming | Technical Considerations |
|---|---|---|---|
| Reprogramming Factors | OCT4, SOX2, KLF4, c-MYC (OSKM) [34] | Core transcription factors for inducing pluripotency | Optimal stoichiometry is critical; c-MYC increases tumorigenic risk |
| Alternative Factors | L-MYC, N-MYC, NANOG, LIN28 [34] | Safer alternatives with similar functionality | L-MYC reduces tumorigenic risk while maintaining efficiency |
| mRNA Modifications | Pseudouridine, 1-methyl pseudouridine [55] | Enhance stability and reduce immunogenicity | Improve translation efficiency and protein yield |
| Delivery Systems | Ionizable LNPs, SORT nanoparticles [55] | mRNA protection and cellular delivery | Influence biodistribution and cell-type specificity |
| Small Molecule Enhancers | Valproic acid, Sodium butyrate, RepSox [1] [34] | Epigenetic modifiers that enhance reprogramming | Can replace some transcription factors (e.g., RepSox replaces SOX2) |
| Characterization Tools | Pluripotency markers (Tra-1-60, SSEA-4) [28] | Validate successful reprogramming | Essential for quality control of resulting iPSCs |
Kinetic Modeling and Pathway Analysis
Computational Approaches to Expression Control
Recent advances in kinetic modeling provide valuable tools for predicting and optimizing protein expression dynamics in reprogramming applications. Ordinary differential equation (ODE) models have been successfully employed to understand how signaling dynamics influence gene expression patterns in related biological systems [56]. These models can be adapted to predict how JNK dynamics—a pathway often involved in reprogramming—contribute to downstream gene expression patterns through regulated transcription factors like c-Jun.
The dynamic encoding principle observed in stress response pathways demonstrates that variations in the temporal dynamics of kinase activation can drive distinct gene expression patterns, partially mediated by mRNA stability [56]. This principle directly applies to reprogramming protocols, where the timing and dynamics of pluripotency factor expression significantly influence the efficiency and quality of resulting iPSCs.
Monitoring and Validation Frameworks
Robust monitoring frameworks are essential for validating precision dosing in reprogramming applications. Recent advances in mass spectrometry methods enable precise measurement of target engagement, providing quantitative data on protein expression levels and function [57]. These bioanalytical methods can be adapted to monitor pluripotency factor expression and function during reprogramming protocols.
For covalent drug development, intact protein mass spectrometry assays have been developed that can analyze drug-target complexes and assess percentage target engagement (%TE) [57]. Similar approaches could be adapted for reprogramming applications to monitor the engagement of pluripotency factors with their genomic targets, providing direct feedback on dosing efficacy.
The precise control of protein expression dynamics through mRNA technology represents a transformative approach in pluripotency research and regenerative medicine. As the field advances, several key areas will shape future progress:
First, the development of increasingly sophisticated delivery systems with enhanced tissue specificity and reduced immunogenicity will improve the precision of reprogramming protocols [55]. Second, the integration of computational modeling and artificial intelligence will enable more accurate predictions of dosing requirements and expression kinetics [58]. Finally, standardized characterization protocols and quality control metrics will ensure consistent outcomes across different research and clinical applications [16].
The convergence of these technologies will accelerate the implementation of mRNA-based reprogramming in both basic research and clinical applications, ultimately enabling robust, reliable production of patient-specific iPSCs for regenerative medicine, disease modeling, and drug discovery. As precision dosing methodologies continue to evolve, mRNA technology is poised to become the gold standard for controlled protein expression in pluripotency research and beyond.
Benchmarking Success: Validating mRNA-iPSCs and Contrasting with Alternative Reprogramming Technologies
The generation of induced pluripotent stem cells (iPSCs) using non-integrating mRNA technology represents a groundbreaking advancement in regenerative medicine, disease modeling, and drug discovery. Unlike earlier methods that relied on viral vectors, mRNA reprogramming avoids genomic integration, significantly reducing the risk of insertional mutagenesis and enhancing the safety profile of derived iPSC lines [1] [22]. This technology involves introducing synthetic mRNA encoding the core transcription factors OCT4, SOX2, KLF4, and c-MYC (OSKM) into somatic cells, effectively reprogramming them to a pluripotent state without altering their genetic code [22].
The non-integrative nature of mRNA-iPSCs makes them particularly valuable for therapeutic applications; however, it also necessitates comprehensive characterization to ensure their quality, stability, and safety. Rigorous validation is essential to confirm that these cells truly exhibit the defining characteristics of pluripotency while maintaining genetic integrity [59]. This whitepaper provides an in-depth technical guide for researchers and drug development professionals, outlining standardized methodologies for evaluating mRNA-iPSCs across three critical domains: pluripotency status, functional differentiation potential, and genetic fidelity. By establishing a robust characterization framework, we can ensure the reliability of mRNA-iPSCs for both basic research and clinical translation.
Morphological Assessment of Putative mRNA-iPSC Colonies
The initial characterization of successfully reprogrammed mRNA-iPSCs begins with a careful morphological assessment under standard culture conditions. This qualitative evaluation serves as a first-line screening tool to identify colonies with characteristic pluripotent features.
Key Morphological Criteria
Undifferentiated human iPSCs grown on feeder layers or in feeder-free conditions exhibit distinctive morphological features that differentiate them from partially reprogrammed or differentiated cells [59]. These characteristics include:
- Colony Architecture: Colonies should be tightly packed, flat-to-multilayered with well-defined, smooth borders, and exhibit a characteristic "spheroidal" appearance when viewed under low magnification [59].
- Cellular Morphology: Individual cells within colonies should appear small and compact with a high nucleus-to-cytoplasm ratio. The cells should display a prominent nucleolus and scant cytoplasm, giving them a "primitive" appearance [59].
- Quantitative Parameters: Wakao et al. established specific quantitative metrics for optimal iPSC morphology, including a single nucleolus, nucleus-to-nucleolus ratio of approximately 2.19, nucleus-to-cytoplasm ratio of about 0.87, cell density of approximately 5,900 cells/mm², and individual cell size around 43.5 μm² [59].
Imaging Methodologies
Phase-contrast microscopy is the most widely used technique for routine morphological evaluation of live iPSC cultures [59]. This method provides enhanced contrast without staining, allowing for continuous monitoring of cell proliferation and colony expansion. For more detailed analysis, fluorescent microscopy can be employed to assess specific cellular structures, while electron microscopy offers ultrastructural resolution but is typically reserved for specialized investigations [59].
Table 1: Microscopy Techniques for Morphological Assessment
| Technique | Applications | Advantages | Limitations |
|---|---|---|---|
| Phase-contrast Microscopy | Routine monitoring of live cells, colony morphology assessment | No staining required, enables time-lapse imaging, cost-effective | Halo artifacts may complicate image analysis |
| Fluorescent Microscopy | Evaluation of specific markers, cell health assessment | High contrast and resolution, specific labeling | Requires staining/fixing, potential phototoxicity |
| Computer-assisted Microscopy | High-content screening, quantitative morphology | Automated analysis, reduced subjectivity, high-throughput | Requires specialized software and validation |
Molecular Characterization of Pluripotency
Molecular characterization provides quantitative validation of the pluripotent state through the detection of specific markers associated with the undifferentiated condition. This analysis occurs at both the protein and gene expression levels.
Surface and Intracellular Marker Analysis
The International Stem Cell Initiative (ISCI) has established a core set of markers for validating pluripotency in human iPSCs. These include specific surface antigens and intracellular transcription factors that are highly expressed in undifferentiated cells [59].
Flow Cytometry represents the gold standard for quantitative analysis of pluripotency markers and is considered a mandatory release criterion for iPSC banking [59]. This technique enables simultaneous detection of multiple markers at the single-cell level, providing statistical data on population homogeneity. The core pluripotency markers to be assessed include:
- Surface Antigens: SSEA3, SSEA4, TRA-1-60, and TRA-1-81 should be highly expressed, while SSEA1 (typically expressed during differentiation) should be absent or minimally detected [59].
- Intracellular Transcription Factors: OCT4, NANOG, and SOX2 represent core pluripotency factors that can be detected after cell permeabilization.
For researchers requiring isolation of specific subpopulations, Fluorescence-Activated Cell Sorting (FACS) extends flow cytometry capabilities by enabling purification of live cells based on marker expression profiles [59].
Gene Expression Analysis
Gene expression analysis complements protein detection by verifying the transcriptional activation of pluripotency networks.
- Quantitative Reverse Transcription PCR (qRT-PCR): This method provides sensitive, quantitative assessment of pluripotency-associated gene expression. Key markers to evaluate include OCT4, NANOG, SOX2, along with other genes characteristic of pluripotent state such as TDGF1, DNMT3B, GABRB3, and GDF3 [59]. Expression levels should be compared to established human embryonic stem cell lines as reference standards.
- Immunocytochemistry: While primarily qualitative, immunocytochemistry offers spatial information about marker distribution within colonies, revealing heterogeneity that might be missed in bulk analyses [59]. This technique is particularly valuable for detecting early signs of spontaneous differentiation through irregular marker expression patterns.
Table 2: Essential Pluripotency Markers for mRNA-iPSC Characterization
| Marker Category | Specific Markers | Expected Expression | Detection Methods |
|---|---|---|---|
| Surface Antigens | SSEA3, SSEA4, TRA-1-60, TRA-1-81 | High expression | Flow cytometry, Immunocytochemistry |
| Differentiation Surface Marker | SSEA1 | Absent/Low expression | Flow cytometry, Immunocytochemistry |
| Transcription Factors | OCT4, NANOG, SOX2 | High expression | qRT-PCR, Flow cytometry (intracellular), Immunocytochemistry |
| Pluripotency-associated Genes | TDGF1, DNMT3B, GABRB3, GDF3 | High expression | qRT-PCR |
Genetic Fidelity and Stability Assessment
Ensuring genetic integrity is particularly crucial for mRNA-iPSCs intended for therapeutic applications. Comprehensive genomic analysis safeguards against reprogramming-induced mutations and culture-acquired abnormalities.
Karyotype Analysis
Standard G-banding karyotyping at a resolution of 400-550 bands remains a mandatory release test for iPSC banking, capable of detecting chromosomal abnormalities larger than 5-10 Mb [59]. This technique provides a genome-wide screen for gross chromosomal rearrangements, aneuploidies, and translocations that might occur during reprogramming or extended culture. Cells should be analyzed at passage numbers relevant to their intended use, with periodic reassessment during long-term culture.
High-Resolution Genomic Analysis
For more sensitive detection of submicroscopic genetic alterations, advanced techniques offer enhanced resolution:
- Comparative Genomic Hybridization (CGH) Array: This method identifies copy number variations (CNVs) down to 10-100 kb resolution, providing a comprehensive profile of chromosomal imbalances that might escape detection by conventional karyotyping [59].
- Single Nucleotide Polymorphism (SNP) Analysis: SNP arrays simultaneously detect copy number variations and loss of heterozygosity (LOH) at high resolution, while also providing genetic fingerprinting for line identification [59].
Recent studies have demonstrated that iPSCs maintain donor-specific epigenetic patterns that are most strongly associated with genetic variation at the iPSC stage, though this relationship may weaken following differentiation [60]. This underscores the importance of comprehensive genetic characterization early in the pipeline.
Functional Characterization of Differentiation Potential
The definitive proof of pluripotency lies in demonstrating the ability to differentiate into derivatives of all three germ layers. Functional assays provide this critical validation through both in vitro and in vivo approaches.
In Vitro Differentiation
The formation of embryoid bodies (EBs) represents a fundamental in vitro method for assessing multi-lineage differentiation potential [59]. When cultured in suspension, iPSCs spontaneously aggregate into three-dimensional EBs that initiate differentiation into cell types representing ectoderm, mesoderm, and endoderm. Following EB formation, differentiation should be confirmed through:
- Immunocytochemical analysis of germ layer-specific markers: β-III-tubulin (ectoderm), α-smooth muscle actin (mesoderm), and α-fetoprotein (endoderm).
- Gene expression profiling via qRT-PCR to quantify marker expression for each germ layer.
- Flow cytometric quantification of differentiated cell populations to assess efficiency and homogeneity.
For more directed differentiation approaches, protocol-specific markers should be employed to verify the generation of target cell types, such as motor neurons or cardiomyocytes [1] [60].
In Vivo Differentiation
The teratoma formation assay represents the most stringent test for pluripotency, though it is typically reserved for research applications due to its complexity [59]. This assay involves injecting iPSCs into immunodeficient mice and allowing tumors to develop over 8-12 weeks. Histological analysis of resulting teratomas should reveal well-differentiated tissues representing all three germ layers, such as:
- Ectoderm: neural epithelium, pigmented cells, stratified squamous epithelium
- Mesoderm: cartilage, bone, muscle, adipose tissue
- Endoderm: respiratory epithelium, intestinal epithelium, glandular structures
The following diagram illustrates the core workflow for the comprehensive characterization of mRNA-iPSCs:
Figure 1: Comprehensive mRNA-iPSC Characterization Workflow. This diagram outlines the sequential validation approach for rigorous assessment of mRNA-derived induced pluripotent stem cells, encompassing morphological, molecular, genetic, and functional analyses.
The Scientist's Toolkit: Essential Research Reagents
Successful characterization of mRNA-iPSCs requires specific reagents and tools designed to assess pluripotency and genetic integrity. The following table details essential solutions for establishing a robust validation pipeline.
Table 3: Research Reagent Solutions for mRNA-iPSC Characterization
| Reagent Category | Specific Examples | Application/Function | Key Considerations |
|---|---|---|---|
| Flow Cytometry Antibodies | Anti-SSEA3, SSEA4, TRA-1-60, TRA-1-81, OCT4, NANOG | Quantitative detection of pluripotency markers at single-cell level | Validate with appropriate isotype controls; check species cross-reactivity |
| qPCR Primers | OCT4, SOX2, NANOG, TDGF1, DNMT3B, GABRB3, GDF3 | Gene expression analysis of pluripotency network | Normalize to multiple housekeeping genes; establish reference expression levels |
| Karyotyping Kits | G-banding solutions, Giemsa stain, Colcemid | Chromosomal analysis for gross abnormalities | Analyze metaphase spreads; minimum 20 metaphases examined |
| Genomic Analysis Arrays | CGH arrays, SNP genotyping arrays | High-resolution detection of CNVs and LOH | Compare to parental somatic cells when possible |
| Differentiation Media | Commercially available trilineage differentiation kits | In vitro assessment of differentiation potential | Include undifferentiated controls; validate with multiple markers per germ layer |
| Immunocytochemistry Antibodies | Germ layer-specific markers (β-III-tubulin, α-SMA, AFP) | Functional validation of differentiation potential | Include appropriate secondary antibodies with minimal cross-reactivity |
Advanced Considerations for mRNA-iPSC Characterization
As mRNA-iPSC technology advances toward clinical applications, additional characterization dimensions warrant consideration:
Epigenetic Memory and Donor-Specific Patterns
Recent evidence indicates that iPSCs maintain donor-specific epigenetic patterns that are most strongly associated with genetic variation at the iPSC stage, though this relationship weakens following differentiation [60]. This epigenetic memory can influence differentiation efficiency and should be monitored through:
- DNA methylation profiling to identify aberrant methylation patterns
- Chromatin accessibility assays (e.g., ATAC-seq) to evaluate epigenetic states
- Histone modification analysis to ensure proper resetting of epigenetic marks
Integration with Gene Editing Technologies
The combination of mRNA reprogramming with CRISPR-based gene editing represents a powerful approach for generating genetically modified iPSCs for disease modeling and therapeutic applications. Recent advances in sequential factor delivery have enabled efficient knock-in of transgenes in clinical-grade iPSCs using virus-free, GMP-compatible methods [61]. When utilizing gene-edited mRNA-iPSCs, additional validation should include:
- Off-target analysis through whole-genome sequencing
- On-target efficiency quantification via digital PCR or next-generation sequencing
- Transgene expression and functionality assessment in differentiated cells
The following diagram illustrates a robust gene editing workflow compatible with clinical-grade mRNA-iPSCs:
Figure 2: GMP-compatible Gene Editing Workflow for mRNA-iPSCs. This sequential factor delivery approach enables efficient knock-in of transgenes without viral vectors or antibiotic selection, maintaining compliance with good manufacturing practice requirements [61].
The comprehensive characterization framework outlined in this technical guide provides a rigorous approach for validating mRNA-iPSCs across morphological, molecular, genetic, and functional domains. By implementing these standardized methodologies, researchers can ensure the quality, safety, and reliability of non-integrating mRNA-derived iPSC lines for both basic research and clinical applications. As the field advances toward therapeutic implementation, continued refinement of characterization standards will be essential for realizing the full potential of this transformative technology in regenerative medicine and drug development.
The generation of induced pluripotent stem cells (iPSCs) represents one of the most significant breakthroughs in regenerative medicine, offering unprecedented potential for disease modeling, drug discovery, and cellular therapeutics. Since the initial discovery that somatic cells could be reprogrammed using defined factors, the field has diversified into multiple technological approaches for delivering reprogramming instructions to target cells. Among these, mRNA-based, viral vector-based, and DNA-based methods have emerged as leading strategies, each with distinct advantages and limitations. This technical guide provides a comprehensive comparison of these three core reprogramming methodologies, framed within the context of a broader thesis on non-integrative mRNA technology for pluripotency research. As the field advances toward clinical applications, understanding the nuanced trade-offs between these platforms becomes increasingly critical for researchers, scientists, and drug development professionals working to translate iPSC technology into safe and effective therapies.
Fundamental Mechanisms
Viral vector-based reprogramming, the original iPSC generation method, primarily utilizes retroviruses or lentiviruses to deliver the canonical Yamanaka factors (OCT4, SOX2, KLF4, and c-MYC) [8] [34]. These systems enable highly efficient transduction and stable genomic integration, ensuring sustained transgene expression throughout the reprogramming process. Lentiviral vectors specifically offer the advantage of transducing non-dividing cells and provide stable, long-term transgene expression, making them valuable research tools [62]. The recent development of non-integrating viral systems, such as Sendai virus, addresses some safety concerns while maintaining high efficiency [16] [1].
DNA-based reprogramming approaches utilize plasmid vectors, including minicircle DNA and episomal plasmids, to deliver reprogramming factors without viral components [1]. These methods typically employ electroporation or lipid-based transfection for intracellular delivery. While DNA vectors avoid the limitations associated with viral packaging capacity, they historically suffered from lower transfection efficiency compared to viral methods. Recent advances in delivery systems, including electroporation devices and lipid nanoparticles, have significantly improved their efficiency [63].
mRNA-based reprogramming represents the newest approach, utilizing synthetic, modified mRNA molecules to encode reprogramming factors [16] [24]. This method delivers genetic instructions directly to the cytoplasm, where they are immediately translated into protein without any risk of genomic integration. The transient nature of mRNA (with a typical half-life ranging from 20 minutes to several hours) requires repeated transfections but eliminates the risk of insertional mutagenesis [63] [24]. Recent innovations in nucleoside modifications and purification methods have reduced the innate immunogenicity of synthetic mRNA, enhancing its utility for clinical applications [24].
Comparative Performance Metrics
Table 1: Key Characteristics of Reprogramming Methods
| Feature | Viral Vector | DNA-Based | mRNA-Based |
|---|---|---|---|
| Genomic Integration | Yes (retro/lentivirus); No (Sendai) | Low frequency | None |
| Reprogramming Efficiency | High (retro/lentivirus: ~0.1%; Sendai: ~0.1-1%) | Moderate (~0.001-0.01%) | High (≥1%) |
| Time to iPSC Generation | 3-4 weeks | 4-6 weeks | 2-3 weeks |
| Safety Profile | Low (integrating); Moderate (non-integrating) | Moderate | High |
| Technical Complexity | Moderate | Moderate to High | High |
| Cost | Moderate | Low | High |
| Regulatory Pathway | Complex | Moderate | Evolving |
| Clinical Translation Potential | Limited (integrating); Promising (non-integrating) | Promising | Highly promising |
Table 2: Molecular and Functional Attributes
| Attribute | Viral Vector | DNA-Based | mRNA-Based |
|---|---|---|---|
| Delivery Target | Nucleus | Nucleus | Cytoplasm |
| Mechanism of Action | Integration (retro/lentivirus) or episomal maintenance (Sendai), then transcription & translation | Nuclear entry, transcription, then translation | Direct translation in cytoplasm |
| Transgene Expression Kinetics | Sustained (weeks-months) | Transient to sustained (days-weeks) | Transient (hours-days) |
| Immunogenicity | Moderate (viral proteins) | Low to Moderate | Moderate to High (can be mitigated with modifications) |
| Footprint-Free iPSCs | No (integrating); Yes (non-integrating) | Yes (with careful screening) | Yes |
| Handling Requirements | BSL-2 typically required | Standard cell culture | Standard cell culture |
Experimental Protocols
mRNA Reprogramming Protocol
Day 0: Plating Somatic Cells
- Isolate and plate human dermal fibroblasts at 5×10^4 cells per well in a 6-well plate using fibroblast medium (DMEM supplemented with 10% FBS, 1% GlutaMAX, and 1% non-essential amino acids)
- Incubate at 37°C with 5% CO2 for 24 hours
Day 1-14: Daily mRNA Transfection
- Prepare lipopolymer or lipid nanoparticle (LNP) complex: Dilute 2 µg of each modified mRNA reprogramming factor (OCT4, SOX2, KLF4, c-MYC, LIN28, and NANOG) in 250 µL of opti-MEM reduced serum medium. In a separate tube, dilute 15 µL of RNAiMAX transfection reagent in 250 µL of opti-MEM. Combine solutions and incubate for 15-20 minutes at room temperature [16] [24]
- Wash cells with DPBS and add 1.5 mL fresh fibroblast medium
- Add mRNA-lipopolymer complexes dropwise to cells while gently swirling plate
- Incubate at 37°C with 5% CO2 for 4-6 hours, then replace with fresh fibroblast medium supplemented with B18R interferon inhibitor (200 ng/mL) to enhance mRNA translation and reduce immune activation
- Repeat transfection daily for 14-21 days
Day 7-21: Medium Transition and Colony Picking
- Around day 7, transition to feeder-free iPSC culture medium (such as mTeSR or E8 medium) when cells show morphological changes indicative of reprogramming
- Monitor emerging iPSC colonies daily
- Pick individual colonies between days 18-28 based on standard iPSC morphology (high nucleus-to-cytoplasm ratio, prominent nucleoli, compact colony formation)
- Transfer picked colonies to Matrigel-coated plates for expansion and characterization
Viral Vector Reprogramming Protocol
Day 0: Viral Transduction
- Plate 1×10^5 human fibroblasts per well in a 6-well plate
- Transduce with lentiviral or Sendai viral vectors encoding OSKM factors at an MOI of 5-10 in the presence of 6 µg/mL polybrene (for lentivirus only)
- Centrifuge plates at 1000 × g for 30 minutes (spinoculation) to enhance transduction efficiency
- Incubate overnight at 37°C
Day 1: Medium Change
- Replace virus-containing medium with fresh fibroblast medium
Day 3-5: Seeding on Feeder Cells
- Trypsinize transduced fibroblasts and re-seed at 5×10^3 to 1×10^4 cells per 10 cm dish containing irradiated mouse embryonic fibroblasts (MEFs) or Matrigel in human ESC medium supplemented with 10 ng/mL bFGF
Day 7-28: Colony Monitoring and Picking
- Change medium daily until iPSC colonies appear (typically 3-4 weeks post-transduction)
- Pick and expand colonies with characteristic iPSC morphology
DNA-Based Reprogramming Protocol
Day 0: Plasmid Transfection
- Plate 1×10^5 human fibroblasts per well in a 6-well plate
- Transfect with oriP/EBNA1-based episomal plasmids (2 µg total DNA) expressing OSKMNL factors using Neon Transfection System (1400V, 20ms, 2 pulses) or lipid-based transfection reagents
- Culture in fibroblast medium for 48 hours
Day 3: Selection and Medium Transition
- Begin selection with appropriate antibiotics if plasmid contains resistance gene
- Transition to human ESC medium 5-7 days post-transfection
- Change medium daily thereafter
Day 21-35: Colony Picking
- Monitor for emerging iPSC colonies
- Pick and expand colonies 3-5 weeks post-transfection
Signaling Pathways and Molecular Mechanisms
Figure 1: mRNA Reprogramming Mechanism - This diagram illustrates the intracellular pathway of mRNA-based reprogramming, highlighting its direct cytoplasmic translation and absence of nuclear steps required by DNA-based methods.
The molecular mechanisms of cellular reprogramming involve complex signaling networks that reset the epigenetic landscape of somatic cells to a pluripotent state. The PI3K/AKT signaling pathway enhances reprogramming efficiency by promoting metabolic switching from oxidative phosphorylation to glycolysis, a hallmark of pluripotent stem cells [16]. Concurrently, TGF-β/SMAD signaling works in concert with WNT signaling to enhance mesenchymal-to-epithelial transition (MET), a crucial early step in reprogramming [1]. The core pluripotency network—comprising OCT4, SOX2, and NANOG—activates a self-reinforcing regulatory circuit that establishes and maintains the pluripotent state through epigenetic modifications including DNA demethylation and histone acetylation [34].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Reprogramming Methods
| Reagent/Category | Specific Examples | Function & Application |
|---|---|---|
| Reprogramming Factors | OCT4, SOX2, KLF4, c-MYC (OSKM); NANOG, LIN28 | Core transcription factors that induce pluripotency; various combinations used across platforms |
| Delivery Vehicles | Lentivirus, Sendai virus, LNPs, Electroporation systems | Enable intracellular delivery of reprogramming factors |
| Culture Media | Fibroblast medium, mTeSR1, E8 medium | Support somatic cell growth and pluripotent stem cell maintenance |
| Small Molecule Enhancers | Valproic acid, Sodium butyrate, RepSox, CHIR99021 | Epigenetic modifiers and signaling pathway agonists that enhance reprogramming efficiency |
| Surface Coatings | Matrigel, Laminin-521, Vitronectin | Provide optimal extracellular matrix for iPSC attachment and growth |
| Characterization Tools | Antibodies against TRA-1-60, SSEA4, OCT4; Pluripotency assays | Validate successful reprogramming and pluripotent state |
The comparative analysis of mRNA, viral vector, and DNA-based reprogramming methods reveals a complex landscape of technological trade-offs. Viral vectors, particularly non-integrating systems like Sendai virus, offer high efficiency and remain valuable research tools but face regulatory hurdles for clinical translation. DNA-based methods provide a non-viral alternative with improving efficiency but still present challenges with delivery efficiency and potential genomic integration. mRNA-based reprogramming emerges as a particularly promising approach for clinical applications due to its non-integrating nature, high efficiency, and precisely controllable expression kinetics. While immunogenicity and delivery optimization remain active areas of investigation, the rapid advancement of mRNA platform technologies, bolstered by recent successes in vaccine development, positions this method as a leading candidate for generating clinical-grade iPSCs. As the field progresses, the optimal choice of reprogramming method will continue to depend on the specific application, with mRNA technology offering a compelling pathway toward safe and effective pluripotent stem cell-based therapies.
The advent of non-integrative reprogramming technologies has revolutionized pluripotency research and regenerative medicine, offering pathways to generate induced pluripotent stem cells (iPSCs) without permanent genetic alterations. Among the most prominent approaches are mRNA-based systems, Sendai virus vectors, and protein-based delivery methods, each presenting distinct technical trade-offs in efficiency, safety, and practical implementation. These non-integrating methods have emerged to overcome the significant safety concerns associated with earlier viral methods that permanently integrated reprogramming factors into the host genome, carrying risks of insertional mutagenesis and oncogene activation [64]. The core challenge in the field lies in balancing reprogramming efficiency against safety profiles and technical complexity, a decision that profoundly impacts research outcomes and clinical translation potential. This analysis provides a structured comparison of these three leading non-integrative technologies, examining their molecular mechanisms, performance metrics, and optimal applications within pluripotency research and therapeutic development.
mRNA-Based Reprogramming
mRNA-based reprogramming utilizes synthetic messenger RNA molecules encoding key pluripotency factors to reprogram somatic cells. The fundamental advantage of this approach lies in its completely synthetic nature and rapid kinetics. The mRNA molecules are engineered with specific modifications to enhance stability and reduce immunogenicity, such as nucleotide substitutions including 5-methylcytosine (m5C) and pseudouridine (Ψ) which help evade cellular pattern recognition receptors [23]. Structurally, these mRNA constructs contain 5' cap structures, optimized 5' and 3' untranslated regions (UTRs), the open reading frame (ORF) encoding reprogramming factors, and a poly(A) tail – all optimized for enhanced translational efficiency and reduced degradation [23] [65]. The mechanism involves repeated transfections of the modified mRNA into target cells, where it is translated in the cytoplasm to produce the reprogramming proteins without any nuclear involvement or genomic integration. However, a significant challenge is the inherent immunogenicity of exogenous RNA, which can trigger antiviral responses and inhibit reprogramming efficiency. Additionally, the requirement for multiple transfections over several days adds technical complexity and potential variability to the process.
Sendai Virus (SeV) Vectors
Sendai virus is an RNA virus belonging to the paramyxovirus family that serves as an efficient delivery vehicle for reprogramming factors. Its key advantage is the cytoplasmic replication without any genomic integration phase, as it remains exclusively in the cell's cytoplasm throughout its life cycle [66] [64]. The virus is engineered to be replication-competent but non-integrating, with temperature-sensitive variants available that can be eliminated through temperature shifting once reprogramming is complete [64]. Standard Sendai virus kits typically contain a combination of vectors expressing the classic Yamanaka factors (OCT4, SOX2, KLF4, with c-MYC often provided separately) [66]. The vectors are also designed with additional safety features such as the absence of pathogenicity in humans and the inability to undergo permanent genomic integration. A comparative study analyzing reprogramming success rates across different methods and source cells found that Sendai virus transduction yielded significantly higher success rates relative to episomal reprogramming methods, making it one of the most efficient non-integrative approaches available [66].
Protein-Induced Pluripotent Stem Cells (piPSCs)
Protein-induced reprogramming represents the most direct approach, involving the delivery of purified recombinant reprogramming proteins into target cells. This method typically utilizes cell-penetrating peptides such as poly-arginine anchors to facilitate the intracellular delivery of the transcription factor proteins [67]. The proteins themselves are often fused to protein transduction domains that enhance cellular uptake and nuclear localization. The primary advantage of this technology is the complete avoidance of genetic manipulation, as no DNA or RNA sequences are introduced into the cell [67]. However, this approach faces significant challenges related to protein stability, the need for repeated applications, and exceptionally low reprogramming efficiency compared to nucleic acid-based methods. Additionally, producing functional, purified transcription factor proteins with proper post-translational modifications presents substantial manufacturing challenges. While this method offers the theoretically safest profile by completely eliminating risks associated with nucleic acid delivery, its practical implementation remains technically demanding with variable outcomes across different cell types.
Comparative Performance Analysis
Table 1: Comparative Analysis of Key Performance Metrics
| Parameter | mRNA-Based | Sendai Virus (SeV) | Protein-Based (piPSCs) |
|---|---|---|---|
| Reprogramming Efficiency | Moderate | High (Significantly higher than episomal methods) [66] | Low |
| Time to iPSC Colonies | ~2-3 weeks | ~3-4 weeks | ~6-8 weeks |
| Genomic Integration Risk | None | None (Cytoplasmic only) [64] | None |
| Immunogenicity | High (Triggers innate immune response) | Moderate (Viral vector) | Low |
| Clearance/Turnover | Rapid degradation (hours) | Can be eliminated with temperature-sensitive strains [64] | Rapid degradation (hours) |
| Technical Complexity | High (Multiple transfections) | Moderate (Single transduction) | High (Repeated protein delivery) |
| Ease of Use | Requires optimization of mRNA design and delivery | Commercially available kits | Technically challenging, low efficiency |
| Safety Profile | High (once immunogenicity managed) | High (non-integrating, removable) | Very High (no genetic material) |
| Cost | Moderate | High | High |
Table 2: Molecular and Mechanistic Characteristics
| Characteristic | mRNA-Based | Sendai Virus (SeV) | Protein-Based (piPSCs) |
|---|---|---|---|
| Reprogramming Factors | Modified mRNA encoding OSKM or OSNL | Viral vectors expressing OSKM | Recombinant OSKM proteins |
| Delivery Mechanism | Transfection (e.g., lipofection) | Viral transduction | Cell-penetrating peptides |
| Location of Action | Cytoplasm | Cytoplasm | Nucleus |
| Persistence | Transient (degrades rapidly) | Persistent until cleared | Transient (degrades rapidly) |
| Key Advantages | Rapid production, definable state, no viral elements | High efficiency, broad cell type applicability, proven protocol | No genetic material, theoretically safest approach |
| Key Limitations | High immunogenicity, requires multiple transfections | Potential persistent infection, immune clearance needed, commercial cost | Extremely low efficiency, protein stability issues, complex production |
Methodological Protocols
mRNA-Based Reprogramming Protocol
The mRNA reprogramming protocol involves daily transfections of modified mRNA over a period of 12-18 days. Critical to success is the careful design of the mRNA constructs to include modified nucleotides (pseudouridine and 5-methylcytosine) that reduce innate immune recognition while maintaining high translational efficiency [23]. The 5' and 3' untranslated regions should be optimized for enhanced stability – commonly used UTRs derive from genes such as alpha-globin. The typical workflow begins with the isolation and plating of human fibroblasts or peripheral blood mononuclear cells (PBMCs) in optimized culture conditions. Daily transfections are performed using lipid-based nanoparticles or other transfection reagents, with the mRNA cocktail containing the reprogramming factors (OCT4, SOX2, KLF4, c-MYC, and optionally LIN28 and NANOG). Between transfections, cells are maintained in specialized media containing innate immune inhibitors to mitigate the response to exogenous RNA. Emerging approaches also incorporate self-amplifying mRNA (saRNA) designs that encode both the antigen of interest and viral replication machinery, enabling longer-lasting protein expression from a lower initial dose [23] [65]. Colonies typically begin to appear within 2-3 weeks and are selected based on embryonic stem cell-like morphology.
Sendai Virus Reprogramming Protocol
The Sendai virus protocol utilizes commercially available kits such as the CytoTune Sendai Reprogramming Kit. The process begins with the plating of target cells (fibroblasts, PBMCs, or other somatic cells) at appropriate densities. After 24 hours, cells are transduced with a combination of SeV vectors, each expressing one of the reprogramming factors (OCT4, SOX2, KLF4, and c-MYC), typically at defined multiplicities of infection (MOI). The transduction medium is replaced after 24 hours with fresh culture medium, and cells are subsequently cultured with regular medium changes every 2-3 days. Transduction efficiency can be monitored through included reporter genes such as EmGFP. Approximately one week post-transduction, transduced cells are harvested and replated onto feeder layers or extracellular matrix-coated plates. The emerging iPSC colonies usually become visible within 3-4 weeks and can be manually picked for expansion and characterization. A critical advantage of this system is the availability of temperature-sensitive SeV strains that can be eliminated by shifting cultures to 39°C after successful reprogramming, providing a method to clear the vector from the resulting iPSCs [64].
Protein-Induced Reprogramming Protocol
The protein-based reprogramming method represents the most technically challenging approach. The protocol involves repeated application of recombinant reprogramming proteins, typically fused to cell-penetrating peptides such as poly-arginine anchors to facilitate cellular uptake [67]. The production of these recombinant proteins requires careful consideration of proper folding, post-translational modifications, and functional activity. The target somatic cells are exposed to the protein cocktail daily or every other day over an extended period of 6-8 weeks, with the proteins needing to reach the nucleus in sufficient quantities to activate the pluripotency network. The low permeability and rapid degradation of the proteins necessitate high concentrations and frequent applications, making the process resource-intensive. Medium is changed regularly, and emerging colonies are monitored for embryonic stem cell-like morphology. Due to the extremely low efficiency of this method (typically <0.001%), careful screening and validation of resulting colonies are essential. The primary advantage remains the complete absence of genetic material, eliminating risks associated with nucleic acid persistence or integration.
Visualization of Reprogramming Pathways
Figure 1: Molecular Pathways of Non-Integrative Reprogramming Approaches. Each method follows distinct molecular routes to activate the pluripotency network without genomic integration.
Research Reagent Solutions Toolkit
Table 3: Essential Research Reagents for Non-Integrative Reprogramming
| Reagent Category | Specific Examples | Function & Application |
|---|---|---|
| Reprogramming Factors | Yamanaka factors (OCT4, SOX2, KLF4, c-MYC), Thomson factors (OCT4, SOX2, NANOG, LIN28) | Core transcription factors that induce pluripotency in somatic cells |
| mRNA Modifications | Pseudouridine (Ψ), 5-methylcytidine (m5C) [23] | Reduce immunogenicity and enhance stability of synthetic mRNA |
| Delivery Systems | Lipid nanoparticles (LNPs), Transfection reagents, Cell-penetrating peptides (CPPs) | Facilitate cellular uptake of nucleic acids or proteins |
| Viral Vectors | CytoTune Sendai Viruses, Temperature-sensitive SeV strains [66] | Efficient delivery of reprogramming factors with cytoplasmic persistence |
| Immune Suppressors | B18R interferon inhibitor, Small molecule inhibitors | Counteract innate immune response against exogenous RNA |
| Culture Media | Pluripotent stem cell media (e.g., mTeSR1), Feeder-free culture systems | Support growth and maintenance of emerging iPSC colonies |
| Characterization Tools | Antibodies against pluripotency markers (OCT4, SOX2, NANOG), Karyotyping kits, Differentiation kits | Validate pluripotency and genetic stability of resulting iPSCs |
The comparative analysis of mRNA, Sendai virus, and protein-based non-integrative approaches reveals a complex landscape of technical trade-offs with no single solution universally superior. The selection of an appropriate method must be guided by specific research objectives, technical capabilities, and safety requirements. mRNA-based technology offers a defined, synthetic system with rapid kinetics but requires careful management of immunogenicity. Sendai virus vectors provide high efficiency and reliability at the cost of viral complexity and potential persistence concerns. Protein-based approaches deliver the ultimate safety profile but with challenging efficiency limitations. For most research applications requiring robust iPSC generation, Sendai virus remains the gold standard balancing efficiency and safety. However, mRNA-based methods show tremendous promise as technology advances in managing immune responses and improving delivery efficiency. The future of pluripotency research will likely see increased refinement of mRNA and protein platforms alongside the development of hybrid approaches that combine the strengths of multiple technologies. As these non-integrative methods continue to evolve, they will progressively enable safer, more efficient cellular reprogramming for both basic research and clinical applications.
The field of regenerative medicine has been fundamentally transformed by the emergence of induced pluripotent stem cells (iPSCs), which offer unprecedented potential for patient-specific therapies without the ethical concerns associated with embryonic stem cells [21] [50]. Among the various reprogramming methods, mRNA-based technology represents a groundbreaking approach for generating clinical-grade iPSCs through non-integrative, transient expression of reprogramming factors [16] [9]. This technology enables precise control over cellular reprogramming while eliminating the risk of genomic integration inherent in early viral vector systems [21] [50]. The core innovation lies in using synthetic mRNA to deliver the essential pluripotency factors – typically OCT4, SOX2, KLF4, and c-MYC (OSKM) – directly into somatic cells, effectively reprogramming them to a pluripotent state without altering their genetic code [16] [34].
The broader thesis of non-integrative mRNA technology positions this approach as a cornerstone for the future of pluripotency research and clinical translation [9]. Unlike traditional gene therapy approaches that permanently modify the host genome, mRNA therapeutics offer a non-integrative and controllable strategy for expressing therapeutic proteins, making them particularly suitable for regenerative applications where safety profiles are paramount [9]. The transient nature of mRNA-mediated expression eliminates the risk of insertional mutagenesis while providing sufficient duration for effective reprogramming, striking an optimal balance between efficacy and safety for clinical applications [16] [50]. As the field advances, mRNA-iPSC technology continues to evolve through improvements in mRNA chemistry, delivery platforms, and manufacturing processes, accelerating its path toward commercial scalability and therapeutic implementation [9].
Technical Foundations and Current Capabilities
Molecular Mechanisms of mRNA Reprogramming
The process of reprogramming somatic cells to pluripotency using mRNA technology involves sophisticated mechanisms of epigenetic remodeling and transcriptional reactivation [50]. When synthetic mRNA encoding the Yamanaka factors is introduced into differentiated cells, it is translated into proteins that initiate a cascade of molecular events. This begins with the suppression of somatic cell identity genes and progresses through the activation of the endogenous pluripotency network [34]. The reprogramming process typically occurs in two distinct phases: an early phase where somatic identity is systematically suppressed, followed by a late phase characterized by stabilization of the pluripotency network [50].
At the molecular level, the mRNA-encoded transcription factors facilitate extensive chromatin remodeling through several mechanisms. The c-MYC component associates with histone acetyltransferase complexes to induce global histone acetylation, which subsequently enables exogenous OCT4 and SOX2 to access their target loci [34]. Simultaneously, epigenetic modifiers such as TET enzymes – enhanced by supplements like vitamin C – promote active DNA demethylation at key regulatory genes including OCT4 [50]. This coordinated epigenetic resetting establishes activating histone marks (H3K4me3) at pluripotency loci while reducing repressive marks (H3K27me3), ultimately creating an open chromatin configuration permissive for establishing pluripotency [50]. The entire process is further modulated by signaling pathways including BMP, Wnt, and TGF-β, which facilitate critical transitions such as the mesenchymal-to-epithelial transition (MET) that is essential for successful reprogramming [50].
Comparative Analysis of Reprogramming Technologies
The table below provides a technical comparison of mRNA-based reprogramming against other established methods, highlighting key parameters relevant to clinical translation:
Table 1: Comparative Analysis of iPSC Reprogramming Technologies
| Reprogramming Method | Genomic Integration | Reprogramming Efficiency | Safety Profile | Clinical Translation Potential | Key Advantages |
|---|---|---|---|---|---|
| mRNA-based | None | Moderate to High | Excellent – no integration, transient expression | High – minimal safety concerns, GMP-compatible | Precise temporal control, no DNA damage risk, suitable for autologous therapy |
| Sendai Virus | None | High | Good – non-integrating but viral persistence | Moderate – viral vector requires clearance | High efficiency, works with difficult-to-transfect cells |
| Episomal Vectors | Very rare | Low to Moderate | Good – predominantly non-integrating | Moderate – potential for plasmid persistence | DNA-based but non-integrating, cost-effective |
| Integrating Viral Vectors | High | High | Poor – insertional mutagenesis risk | Low – significant safety concerns | High efficiency, well-established protocol |
| Protein Transduction | None | Very Low | Excellent – completely non-genetic | Low – technically challenging, inefficient | No genetic material introduced, theoretically safest |
The superior safety profile of mRNA technology stems from its completely DNA-free approach, which eliminates not only integration risks but also potential epigenetic aberrations associated with vector-based methods [16] [50]. Additionally, mRNA reprogramming demonstrates faster kinetics compared to other non-integrating methods, with colony emergence typically occurring within 2-3 weeks post-transfection [50]. The technology also offers the unique advantage of precise control over reprogramming factor stoichiometry through careful mRNA design and dosing, enabling optimization of reprogramming efficiency and quality of resulting iPSC colonies [34].
Key Workflow and Signaling Pathways
The mRNA reprogramming process follows a defined workflow with critical signaling pathways activated at specific stages. The diagram below illustrates the core experimental workflow and the key signaling pathways involved in successful cellular reprogramming:
Diagram 1: mRNA Reprogramming Workflow and Pathways
The workflow initiates with the isolation of somatic cells, typically dermal fibroblasts or peripheral blood mononuclear cells (PBMCs), which are then subjected to repeated transfections with synthetic mRNA encoding the reprogramming factors [50] [34]. Critical signaling pathways are activated sequentially throughout the process: the Wnt/β-catenin pathway facilitates the initial metabolic shift toward glycolysis; TGF-β/SMAD signaling promotes the mesenchymal-to-epithelial transition (MET); while PI3K/AKT and MAPK/ERK pathways support cell survival and proliferation during colony formation [50] [34]. This coordinated activation of multiple signaling cascades enables the extensive epigenetic remodeling necessary for establishing pluripotency, ultimately yielding iPSC colonies that can be isolated, expanded, and rigorously characterized for downstream applications.
Clinical Translation Landscape
Current Clinical Trials and Applications
The clinical translation of mRNA-iPSC technology has gained significant momentum, with multiple therapeutic applications now in various stages of clinical development. The table below summarizes key clinical applications and their current development status:
Table 2: Clinical Pipeline for iPSC-Derived Therapies
| Therapeutic Area | Target Condition | Cell Type | Development Stage | Key Organizations/ Trials |
|---|---|---|---|---|
| Neurodegenerative Diseases | Parkinson's Disease | Dopaminergic neurons | Phase I/II trials | Sawamoto et al. 2025; Mass General Brigham autologous trial |
| Ocular Disorders | Age-related Macular Degeneration | Retinal pigment epithelial (RPE) cells | IND approval (2024); Clinical trials | Eyecyte-RPE (India); Healios K.K. |
| Cardiovascular Diseases | Heart Failure | Cardiomyocytes | Preclinical/Phase I | Heartseed Inc.; Cuorips Inc.; Avery Therapeutics |
| Metabolic Disorders | Diabetes | Pancreatic beta cells | Preclinical development | Allele Biotechnology |
| Musculoskeletal Disorders | Osteoarthritis | Chondrocytes | Preclinical/early clinical | Mayo Clinic program |
| Oncology | Hematologic malignancies | iPSC-derived NK cells, CAR-T cells | Multiple Phase I/II | Fate Therapeutics; Century Therapeutics; Editas Medicine |
Recent clinical milestones demonstrate tangible progress in the field. A Phase I/II trial published in April 2025 reported that allogeneic iPSC-derived dopaminergic progenitors successfully survived transplantation, produced dopamine, and did not form tumors in Parkinson's patients [21] [50]. Concurrently, an ongoing autologous iPSC-derived dopamine neuron trial at Mass General Brigham is pioneering the use of a patient's own blood-derived iPSCs for Parkinson's disease, eliminating the need for immune suppression [21] [50]. In the retinal field, Eyecyte-RPE, an iPSC-derived retinal pigment epithelium product, received IND approval in India in 2024 for geographic atrophy associated with age-related macular degeneration, representing a significant step toward scalable and cost-effective cell therapy approaches [21] [50].
Assessment of Clinical Readiness
The clinical readiness of mRNA-iPSC technology varies substantially across therapeutic areas, with ophthalmology and neurology currently leading the translation pathway. Several factors contribute to this differential readiness:
Ophthalmologic Applications: Cell therapies for retinal diseases represent the most advanced clinical applications, owing to the immune-privileged status of the eye, the need for relatively modest cell numbers, and the ability to monitor transplanted cells non-invasively [21]. The first iPSC-based therapy for age-related macular degeneration was initiated over a decade ago, providing substantial clinical experience with retinal cell transplantation [34].
Neurological Applications: Parkinson's disease treatments have advanced to Phase I/II trials, benefiting from extensive preclinical work in non-human primates and the relatively defined pathology involving dopaminergic neuron loss [21] [50]. The recent clinical data demonstrating graft survival and absence of tumor formation represents a critical safety milestone for the field [21] [50].
Cardiovascular Applications: iPSC-derived cardiomyocytes for heart failure have shown promise in preclinical models, with companies like Heartseed Inc. and Cuorips Inc. advancing toward clinical trials [68]. However, challenges remain regarding electrical integration of transplanted cells, as evidenced by transient arrhythmias observed in non-human primate studies [21] [50].
Metabolic and Orthopedic Applications: These applications generally remain at earlier developmental stages, though promising preclinical data supports continued investment and development [68] [41].
Across all applications, the transition from autologous to allogeneic approaches using HLA-matched iPSC banks represents a significant trend in clinical translation strategy. Initiatives like the Kyoto University iPSC Research and Application Center are developing comprehensive iPSC banks where 75 carefully selected lines could theoretically cover 80% of the Japanese population through HLA matching [34]. This approach addresses the significant cost and manufacturing challenges associated with patient-specific therapies while still minimizing immune rejection risks.
Manufacturing and Scale-Up Considerations
Current Manufacturing Challenges
The path to commercial-scale manufacturing of mRNA-iPSC therapies presents multiple technical and operational challenges that must be addressed for successful clinical translation. Key limitations include:
Reprogramming Efficiency: Despite improvements, mRNA reprogramming efficiency remains variable across different cell types and donor ages, potentially requiring protocol optimization for specific applications [34]. Current efficiencies typically range from 0.1% to 1%, necessitating robust screening processes to identify high-quality iPSC clones [50] [34].
Process Standardization: The transition from research-grade to clinical-grade iPSC manufacturing requires strict adherence to Good Manufacturing Practice (GMP) standards, including defined, xeno-free culture conditions and comprehensive quality control measures [21] [50]. This standardization is particularly challenging for the mRNA reprogramming process, which involves multiple transfection steps and sensitive timing [50].
Analytical Characterization: Rigorous assessment of iPSC quality including genomic stability, pluripotency verification, and differentiation potential remains time-consuming and requires specialized expertise [21] [34]. The field still lacks universally standardized potency assays for many iPSC-derived cell products, creating regulatory challenges [21].
Scale-Up Limitations: Traditional 2D culture systems for iPSC expansion have limited scalability and are labor-intensive, creating bottlenecks in producing sufficient quantities for widespread clinical use [21] [41]. Transitioning to 3D bioreactor systems presents its own challenges in monitoring and controlling differentiation in three-dimensional environments [68].
Emerging Solutions for Commercial Scale-Up
Several technological innovations are addressing these manufacturing challenges and enabling more robust, scalable production processes:
Automated Production Systems: Companies are developing integrated closed-system automated platforms for iPSC manufacturing and differentiation. For instance, Cellular Origins' Constellation platform incorporates robotic systems for automated cell culture, while 3P innovation's cryoFIL system enables automated fill-finish processes critical for final product formulation [69].
Advanced Bioreactor Technologies: The transition from 2D culture to 3D bioreactor systems enables more efficient expansion of iPSCs and their derivatives [68]. Companies like Cellistic are leveraging 3D bioreactor-based manufacturing platforms for producing iPSC-derived CAR-NKT cells at scales relevant for clinical trials [68].
Process Analytical Technologies: Implementation of advanced monitoring systems including AI-guided image analysis for colony selection and in-line sensors for metabolic parameters enables real-time quality assessment and process control [16] [41]. Machine learning algorithms, such as those described by Vedeneeva et al., can automatically detect and characterize iPSC colonies, enhancing selection of high-quality clones for expansion [16] [21].
Supply Chain Innovations: Integrated cold-chain solutions and distributed manufacturing networks are being developed to support global distribution of iPSC-based therapies [70] [69]. Companies like I Peace, Inc. have implemented fully-closed automated iPSC manufacturing systems that meet stringent regulatory standards while enabling mass production of clinical-grade iPSC lines [68].
The growing pipeline of iPSC-derived therapies has catalyzed significant investment in manufacturing infrastructure. The global induced pluripotent stem cells market size was valued at USD 1.93 billion in 2024 and is predicted to reach approximately USD 5.12 billion by 2034, expanding at a CAGR of 10.25% [41]. This growth is driving innovation in manufacturing technologies and economies of scale that will be critical for making iPSC therapies commercially viable.
Technical Protocols and Research Toolkit
Essential Research Reagents and Materials
The successful implementation of mRNA-iPSC technology requires carefully selected reagents and materials optimized for efficiency, reproducibility, and clinical compatibility. The table below details key components of the research toolkit for mRNA reprogramming:
Table 3: Essential Research Reagent Solutions for mRNA Reprogramming
| Reagent Category | Specific Examples | Function | Clinical Translation Considerations |
|---|---|---|---|
| mRNA Reprogramming Factors | OCT4, SOX2, KLF4, c-MYC (OSKM) synthetic mRNA | Induction of pluripotency through transient expression of key transcription factors | GMP-grade manufacturing; modified nucleotides to reduce immunogenicity |
| mRNA Modifications | Pseudouridine, 5-methylcytidine | Enhanced stability and reduced innate immune recognition | Critical for minimizing cell death and improving reprogramming efficiency |
| Transfection Reagents | Lipid nanoparticles (LNPs), commercial transfection reagents | Delivery of mRNA molecules across cell membrane | Clinical-grade materials required; LNP technology enables high efficiency |
| Cell Culture Media | Defined, xeno-free maintenance media | Support iPSC growth and maintenance under defined conditions | Essential for clinical compliance; eliminates animal-derived components |
| Supplemental Small Molecules | CHIR99021 (GSK3β inhibitor), valproic acid (HDAC inhibitor) | Enhance reprogramming efficiency through epigenetic modulation | Replace viral factors; improve consistency across cell types |
| Characterization Antibodies | Anti-OCT4, SOX2, NANOG, TRA-1-60, SSEA4 | Validation of pluripotency marker expression | Standardized panels for quality control; flow cytometry and immunocytochemistry |
| Differentiation Media | Defined kits for ectoderm, mesoderm, endoderm lineages | Functional validation of pluripotency through trilineage differentiation | Standardized protocols for consistent assessment of iPSC quality |
The selection of appropriate mRNA modifications represents a critical factor in reprogramming success. Incorporation of modified nucleotides such as pseudouridine and 5-methylcytidine significantly reduces activation of innate immune responses while extending mRNA half-life, both of which contribute to improved reprogramming efficiency [9] [50]. Additionally, the use of small molecule supplements like CHIR99021 (a GSK3β inhibitor) and valproic acid (a histone deacetylase inhibitor) has been shown to improve reprogramming efficiency by modulating signaling pathways and epigenetic barriers that would otherwise limit conversion to pluripotency [50].
Detailed Experimental Protocol
A robust protocol for mRNA-mediated reprogramming requires careful attention to multiple critical parameters. The following methodology has been optimized based on current best practices:
Initial Cell Preparation:
- Source somatic cells (typically dermal fibroblasts or PBMCs) from approved donors with appropriate consent
- Culture cells in optimized growth media until 70-80% confluent, ensuring logarithmic growth phase
- Passage cells at least twice after thawing to ensure recovery and stability before reprogramming initiation
mRNA Transfection Protocol:
- Day 0: Plate 50,000-100,000 somatic cells per well in 6-well plates coated with appropriate matrix
- Day 1: Begin daily transfections with mRNA cocktail containing OSKM factors (typically 0.5-1 μg per factor per well)
- Use clinical-grade transfection reagent per manufacturer's instructions
- Include modified nucleotides (pseudouridine, 5-methylcytidine) to reduce immune activation
- Continue daily transfections for 12-16 days, with medium changes 4-6 hours post-transfection
Colony Selection and Expansion:
- Day 18-21: Identify emerging iPSC colonies based on characteristic morphology (compact cells, defined edges, high nucleus-to-cytoplasm ratio)
- Mechanically pick individual colonies or use cell sorting for TRA-1-60 positive cells
- Expand selected colonies in defined, xeno-free culture conditions on appropriate matrices
- Cryopreserve early passage stocks for long-term storage
Quality Control Assessment:
- Perform comprehensive characterization including:
- Pluripotency marker analysis (OCT4, SOX2, NANOG, SSEA4, TRA-1-60) via immunocytochemistry and flow cytometry
- Trilineage differentiation potential through directed differentiation protocols
- Karyotyping and genomic stability assessment (G-band karyotyping or SNP analysis)
- Identity verification through Short Tandem Repeat (STR) profiling
- Endogenous pluripotency gene activation confirmation via RT-PCR
This protocol typically achieves reprogramming efficiencies of 0.2-1.0%, with higher efficiencies observed in younger donor cells and when supplemented with small molecule enhancers [50] [34]. The critical success factors include mRNA quality, precise timing of transfections, careful colony selection, and rigorous quality control throughout the process.
Emerging Technological Innovations
The future clinical translation of mRNA-iPSC technology will be shaped by several emerging innovations that address current limitations:
Advanced Gene Editing Integration: The combination of mRNA reprogramming with CRISPR-Cas9 gene editing enables correction of genetic defects in patient-specific iPSCs before differentiation and transplantation [16] [21]. Newer CRISPR systems including base editors and prime editors allow more precise genetic correction without double-strand breaks, enhancing safety profiles for clinical applications [16].
Immune Evasion Strategies: Research teams are employing CRISPR-Cas9 to engineer hypoimmunogenic iPSCs by deleting HLA class I and II molecules while adding immune regulatory proteins like PD-L1 [16]. This approach aims to create "universal" iPSC lines that evade immune rejection without requiring matching or immunosuppression [16] [21].
Organoid and Tissue Engineering: The creation of 3D organoid models from iPSCs provides more physiologically relevant systems for disease modeling and drug screening [16]. These complex tissue-like structures better recapitulate native tissue architecture and function compared to traditional 2D cultures, enabling more predictive preclinical assessment [16].
AI-Enabled Manufacturing: Artificial intelligence and machine learning are being applied to optimize reprogramming protocols, predict differentiation outcomes, and automate quality control processes [16] [41]. These technologies enhance standardization and reproducibility while reducing manual labor requirements in iPSC manufacturing [16] [21].
Commercialization Outlook
The commercialization pathway for mRNA-iPSC therapies continues to accelerate, with the global market projected to grow at a CAGR of 10.25% from 2025 to 2034 [41]. This growth is fueled by increasing demand for patient-specific cell therapies, advancements in reprogramming technologies, and expanding applications in regenerative medicine [41]. North America currently dominates the global iPSC market with an estimated 45% market share in 2024, though the Asia-Pacific region is expected to grow at the fastest rate during the forecast period, driven by significant government investments and streamlined regulatory frameworks [41].
The regulatory landscape for iPSC-based therapies continues to evolve, with agencies providing increasingly clear pathways for clinical approval [21]. However, regulatory complexity remains a challenge, particularly for therapies with novel mechanisms of action or combined gene editing components [41]. The high costs associated with iPSC development and manufacturing under GMP conditions also present barriers to entry, particularly for smaller organizations [41]. Despite these challenges, the ongoing expansion of HLA-matched iPSC banks and continued technological innovations in manufacturing are expected to progressively reduce costs and increase accessibility of iPSC-based therapies.
In conclusion, mRNA-iPSC technology represents a transformative approach for regenerative medicine, offering an optimal balance of efficacy and safety for clinical applications. While challenges remain in manufacturing scale-up and standardization, the field has demonstrated substantial progress with multiple therapies now in clinical trials. The continued integration of advances in gene editing, bioengineering, and computational technologies will further enhance the development of safe and effective iPSC-based therapeutic options, ultimately enabling broad implementation of personalized regenerative treatments.
Conclusion
Non-integrative mRNA technology has firmly established itself as a cornerstone for the safe and efficient production of clinical-grade iPSCs, effectively addressing the critical safety concern of insertional mutagenesis associated with earlier viral methods. The synthesis of insights from foundational mechanisms to clinical applications confirms that mRNA reprogramming is a robust and versatile platform. However, the path to widespread clinical adoption requires continued optimization to overcome challenges in reprogramming efficiency, immune response control, and scalable manufacturing. Future progress hinges on the convergence of mRNA technology with advancements in bioengineering, such as refined lipid nanoparticles for delivery, and computational tools like AI for quality control. This synergy will accelerate the development of personalized regenerative therapies, on-demand disease models, and innovative cell-based drugs, ultimately unlocking the full potential of iPSC technology in biomedicine.
References
- 1. Advances in Induced Pluripotent Stem Cell Reprogramming ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12577567/]
- 2. Application of the Yamanaka Transcription Factors Oct4 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10531188/]
- 3. mRNA-Based Genetic Reprogramming - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6453511/]
- 4. Generation of iPSCs by Nonintegrative RNA-Based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6607707/]
- 5. Cell Reprogramming and Differentiation Utilizing ... [https://pubmed.ncbi.nlm.nih.gov/38535481/]
- 6. HSCI researchers achieve major breakthrough in cell ... [https://www.hsci.harvard.edu/hsci-researchers-achieve-major-breakthrough-cell-reprogramming]
- 7. Generation of human induced pluripotent stem cells using ... [https://www.sciencedirect.com/science/article/pii/S187350611630006X]
- 8. Current advances and future prospects of cell ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2025.1546423/full]
- 9. mRNA-based technology for engineered regenerative ... [https://www.sciencedirect.com/science/article/pii/S305056232500176X]
- 10. The long and winding road of reprogramming-induced ... [https://www.nature.com/articles/s41467-024-46020-5]
- 11. Induced pluripotent stem cells (iPSCs) [https://www.nature.com/articles/s41392-024-01809-0]
- 12. Transient non-integrative expression of nuclear ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7093390/]
- 13. Transient naive reprogramming corrects hiPS cells ... [https://www.nature.com/articles/s41586-023-06424-7]
- 14. Intercellular mRNA transfer alters the human pluripotent stem ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11789055/]
- 15. A comparison of non-integrating reprogramming methods [https://pmc.ncbi.nlm.nih.gov/articles/PMC4329913/]
- 16. Pluripotent Stem Cells: Recent Advances and Emerging ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12025069/]
- 17. Recombinant Sendai Virus Vectors as Novel Vaccine ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12115985/]
- 18. A Sendai virus-based expression system directs efficient ... [https://www.nature.com/articles/s41598-024-77508-1]
- 19. Deep generative optimization of mRNA codon sequences ... [https://www.nature.com/articles/s41467-025-64894-x]
- 20. Novel centromeric plasmid for stable extrachromosomal ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12227449/]
- 21. Clinical translation of human iPSC technologies: advances ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2025.1627149/full]
- 22. Recent progress in the use of induced pluripotent stem ... [https://link.springer.com/article/10.1007/s44340-025-00042-x]
- 23. Progress and prospects of mRNA-based drugs in pre- ... [https://www.nature.com/articles/s41392-024-02002-z]
- 24. mRNA in Cell and Gene Therapy: From Vaccines to In Vivo ... [https://www.hyperec.com/blog/mrna-in-cell-and-gene-therapy-from-vaccines-to-in-vivo-reprogramming/]
- 25. Potential health risks of mRNA-based vaccine therapy [https://www.sciencedirect.com/science/article/abs/pii/S0306987723000117]
- 26. Cell reprogramming: methods, mechanisms and applications [https://cellregeneration.springeropen.com/articles/10.1186/s13619-025-00229-x]
- 27. High-efficiency RNA-based reprogramming of human ... [https://www.nature.com/articles/s41467-018-03190-3]
- 28. mRNA Cellular Reprogramming Method [https://pluristyx.com/cellular-reprogramming/]
- 29. Current Challenges of iPSC-Based Disease Modeling and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6562607/]
- 30. Quality control guidelines for clinical-grade human induced ... [https://pubmed.ncbi.nlm.nih.gov/30205750/]
- 31. Generation of clinical-grade human induced pluripotent stem ... [https://stemcellres.biomedcentral.com/articles/10.1186/s13287-015-0206-y]
- 32. Manufacturing Parameters for the Creation of Clinical ... [https://pubmed.ncbi.nlm.nih.gov/38402590/]
- 33. GMP iPSC Production Service: Clinical-grade Human ... [https://www.reprocell.com/clinical-stem-cell-services/gmp-ipsc-production]
- 34. A comprehensive analysis of induced pluripotent stem cell ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2025.1593207/full]
- 35. Review mRNA-Based Genetic Reprogramming [https://www.sciencedirect.com/science/article/pii/S1525001618305938]
- 36. A molecular roadmap of cellular reprogramming into iPS cells [https://pmc.ncbi.nlm.nih.gov/articles/PMC3608203/]
- 37. Induced Pluripotent Stem Cells as a Disease Modeling and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3343213/]
- 38. Exploring the promising potential of induced pluripotent ... [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-023-01873-0]
- 39. iPSCs as a Platform for Disease Modeling, Drug Screening ... [https://www.mdpi.com/2073-4409/8/1/20]
- 40. Potential use of iPSCs for disease modeling, drug screening ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10687886/]
- 41. Induced Pluripotent Stem Cells Market Size 2025 to 2034 [https://www.precedenceresearch.com/induced-pluripotent-stem-cells-market]
- 42. mRNA – A game changer in regenerative medicine, cell ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9418126/]
- 43. Highly efficient reprogramming to pluripotency and directed ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3656821/]
- 44. RNA-Based Strategies for Cell Reprogramming toward ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8876983/]
- 45. Innate immune mechanisms of mRNA vaccines - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9641982/]
- 46. Nucleotide Modifications Decrease Innate Immune Response ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6266926/]
- 47. Nucleotide Modifications Decrease Innate Immune ... [https://pubmed.ncbi.nlm.nih.gov/30400232/]
- 48. Innate immune responses against mRNA vaccine promote ... [https://www.nature.com/articles/s41467-024-51411-9]
- 49. Immunosuppressants: Definition, Uses & Side Effects [https://my.clevelandclinic.org/health/treatments/10418-immunosuppressants]
- 50. Clinical translation of human iPSC technologies [https://pmc.ncbi.nlm.nih.gov/articles/PMC12498231/]
- 51. Reassessment of marker genes in human induced ... [https://www.nature.com/articles/s41467-024-52922-1]
- 52. qPCR Analysis for Pluripotency Markers for iPSC [https://www.creative-biolabs.com/stem-cell-therapy/qpcr-analysis-for-pluripotency-markers-for-ipsc.htm]
- 53. Engineered iPSC‐derived natural killer cells [https://pmc.ncbi.nlm.nih.gov/articles/PMC12246386/]
- 54. Error-corrected next-generation sequencing mutagenicity ... [https://www.frontiersin.org/journals/toxicology/articles/10.3389/ftox.2025.1657189/full]
- 55. mRNA therapeutics beyond vaccines: dosing precision ... [https://pubs.rsc.org/en/content/articlehtml/2026/pm/d5pm00159e]
- 56. JNK activation dynamics drive distinct gene expression ... [https://www.nature.com/articles/s41540-025-00590-2]
- 57. Mass spectrometry methods and mathematical PK/PD ... [https://www.nature.com/articles/s41467-025-56985-6]
- 58. From Data to Dose: Diversity in Therapeutic Drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12388870/]
- 59. Methods for Pluripotent Stem Cell Characterization [https://pmc.ncbi.nlm.nih.gov/articles/PMC11815987/]
- 60. iPSCs and iPSC-derived cells as a model of human ... [https://www.nature.com/articles/s41467-025-56569-4]
- 61. Sequential factor delivery enables efficient workflow for ... [https://www.nature.com/articles/s41598-025-17876-4]
- 62. Viral Vector Production (Research-use) Market [https://www.factmr.com/report/viral-vector-production-research-use-market]
- 63. DNA Vaccines in the Post-mRNA Era - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC12429263/]
- 64. Non-integrating Methods to Produce Induced Pluripotent ... [https://www.intechopen.com/chapters/74358]
- 65. Recent Advancement in mRNA Vaccine Development and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10384963/]
- 66. Human iPSC Reprogramming Success: The Impact of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11756937/]
- 67. Advantages & Disadvantages of Induced Pluripotent Stem ... [https://bioinformant.com/induced-pluripotent-stem-cell-ipsc-market/]
- 68. The Pipeline for of iPSC-Derived Cell Therapeutics in 2026 [https://bioinformant.com/ipsc-derived-cell-therapeutics/]
- 69. Industry updates from the field of stem cell research and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11881883/]
- 70. Anemocyte to attend Advanced Therapies Usa 2025 - News [https://anemocyte.com/anemocyte-to-attend-advanced-therapies-usa-2025/]